Thank you for visiting nature.com. You are using a browser version with limited support for CSS. To obtain the best experience, we recommend you use a more up to date browser (or turn off compatibility mode in Internet Explorer). In the meantime, to ensure continued support, we are displaying the site without styles and JavaScript.
- View all journals
- Explore content
- About the journal
- Publish with us
- Sign up for alerts
- Open access
- Published: 14 December 2023

Does students’ awareness of school-track-related stereotypes exacerbate inequalities in education?
- Lisa Bardach ORCID: orcid.org/0000-0002-2168-3117 1 ,
- Claudia Neuendorf ORCID: orcid.org/0000-0002-3024-0000 1 , 2 ,
- Kou Murayama ORCID: orcid.org/0000-0003-2902-9600 1 ,
- Thorsten Fahrbach 1 ,
- Michel Knigge 3 ,
- Benjamin Nagengast ORCID: orcid.org/0000-0001-9868-8322 1 , 4 &
- Ulrich Trautwein ORCID: orcid.org/0000-0003-0647-0057 1
npj Science of Learning volume 8 , Article number: 59 ( 2023 ) Cite this article
1600 Accesses
2 Citations
3 Altmetric
Metrics details
- Human behaviour
Early ability tracking increases inequalities in education. It has been proposed that the awareness of negative school-track-related stereotypes contributes to educational inequalities, as stereotype awareness interferes with students’ abilities to thrive, particularly those in lower, stigmatized tracks. The present study tested this assumption in a sample of 3880 German secondary school students from three tracks, who were assessed four times on stereotype awareness regarding their own school track and academic outcomes (achievement, engagement, self-concept) between Grades 5 and 8. Students in the lowest track reported higher levels of stereotype awareness than higher track students or students attending a combined track. Stereotype awareness increased across time in all tracks. Contrary to our preregistered hypotheses, however, the results from multigroup models revealed that (changes in) stereotype awareness were not more strongly related to (changes in) most outcomes in the lowest track in comparison with the other two tracks.
Similar content being viewed by others
Self-perceptions as mechanisms of achievement inequality: evidence across 70 countries
Simple questionnaires outperform behavioral tasks to measure socio-emotional skills in students
Race, academic achievement and the issue of inequitable motivational payoff
Introduction.
Many education systems all around the globe group students by ability. Often, the sorting of students into different educational tracks is viewed as a way to help educators target students’ learning needs more effectively. At the same time, tracking has lasting consequences for students’ learning and later careers. For instance, educational tracks represent differential developmental contexts, with higher average levels of teaching quality and learning rates in higher tracks 1 , 2 . Further, the assignment to different school tracks has been criticized for reproducing existing social class differences 2 , 3 , as children from less socioeconomically advantaged families have a higher chance of being enrolled in a lower track secondary school. These tracking decisions cannot be explained by lower abilities of disadvantaged groups alone, as it has been shown that teachers provide higher tracking recommendations for students coming from higher socioeconomic status backgrounds than for equally performing students with lower socioeconomic status 4 .
In addition, students’ awareness of stereotypes relating to their school track could further exacerbate inequalities. Stereotypes capture oversimplified beliefs about the characteristics of members of certain groups 5 , and educational tracks generate different stereotypes about the students attending these tracks. Stereotypes regarding students in higher ability tracks (higher status) involve the characterizations that they are “smart” and “perform well,” whereas the opposite is expected from students attending lower ability tracks (lower status) for whom negative stereotypes prevail (e.g., being “stupid or “unmotivated” 3 , 5 , 6 , 7 ). Being aware of negative stereotypes can lead to disengagement from the stigmatized domain and lower domain-specific performance 7 , 8 . Hence, awareness of negative school-track-related stereotypes may become a psychological barrier that particularly hinders the thriving of lower track students.
However, prior quantitative research on students’ awareness of school-track-related stereotypes has been surprisingly scarce 3 , 7 , 9 , leaving serious gaps in the current understanding. The present longitudinal study therefore set out to investigate (a) mean-level differences in students’ awareness of negative school-track-related stereotypes (comprising negative cognitive, motivational, and social stereotypes) between tracks, (b) developmental trajectories of stereotype awareness, and (c) relationships between (changes in) stereotype awareness and (changes in) academic outcomes in terms of academic achievement (standardized achievement test scores), self-concept of academic aptitude, and school engagement. We relied on a sample of German secondary school students from three different nonacademic tracks who were assessed four times on their awareness of stereotypes and all outcomes between the ages of 11 and 14 years (Grades 5, 6, 7, and 8). Specifically, our sample included students from the German federal state of Baden-Württemberg attending Hauptschule, the lowest track in Germany, and Realschule, a higher track. Our sample also included students from Mittelschule, a combined track (students in this track could get a diploma equivalent to Hauptschule or a diploma equivalent to Realschule) from the German federal state of Saxony. Multigroup models were used to examine potentially differentiated patterns of effects in different school tracks. Throughout the manuscript, students’ awareness of school-track-related stereotypes refers to stereotypes pertaining to a student’s own school track (i.e., students from a particular track report how aware they are of stereotypes that pertain only to their track) and not stereotypes that pertain to other tracks.
Does students’ awareness of school-track-related stereotypes differ between tracks ( Research Question 1, RQ1 )? The placement in a relatively lower track (compared with a relatively higher track) can be perceived as a devalued social position reflective of a student’s ability and includes information about the student’s standing in society 10 , 11 , 12 . Tracking thus provides students with institutionalized status labels that are highly visible and impactful (e.g., with respect to later career chances and life paths 9 , 13 ). Moreover, students know about the image of their track in society. Consequently, lower track students are aware of the negative stereotypes that “others” or “people in general” hold about their school track 7 , 9 , even though they do not necessarily endorse these negative stereotypes themselves (see also research on social stigma 14 ).
Differences in stereotype awareness between tracks, with higher levels of stereotype awareness in lower tracks, as well as the content of stereotypes can also be linked to differentiation-polarization theory 10 , 15 , 16 . The theory states that the differentiation of students into different tracks leads to a polarization of the students’ school attitudes. For higher track students, school is a positive experience, given that belonging to a higher track reflects a higher status. By contrast, lower track students lose status due to their assignment to a lower and less valued track. Lower track students therefore react against this system and the values it upholds, namely, ability and hard work. Consequently, an “antischool culture” emerges in the lower tracks 10 , 12 , 15 . Here, we argue that polarization might be reflected not only in school cultures and the corresponding school attitudes (as in the initial formulation of the theory) but also in students’ awareness of how the public perceives their track and the characteristics of students from their track, with negative characteristics that are detrimental to academic success (akin to antischool cultures and attitudes) being attributed to lower track students. Hence, regarding the preregistered RQ1 (differences between tracks), we hypothesized that we would find mean differences in stereotype awareness between tracks in all four waves, with higher average levels of negative school-track-related stereotype awareness for students from the lowest track in comparison with students from the combined track and the higher track.
How does students’ awareness of school-track-related stereotypes develop over time in different tracks ( Research Question 2, RQ2 )? Conceptually, stereotypes have often been considered to be fixed, persisting even in the face of conflicting evidence 17 . Thus, students’ awareness of school-track-related stereotypes could remain stable over time, reflecting stable status differences between tracks within a society that are visible to students. On the other hand, it has been highlighted that stereotypes can undergo developmental changes during the school years 18 , 19 . For students’ awareness of school-track-related stereotypes, both increasing and decreasing trajectories seem theoretically plausible.
Students’ awareness of negative school-track-related stereotypes may increase over time, at least in the lower tracks. During adolescence, students gain a more nuanced understanding of their own social position and become more sensitive to cues reflecting the devalued position of their group (here: lower track students) in society 5 , 20 . These developmental processes likely provide a fertile foundation for lower track students’ stereotype awareness. Alternatively, stereotype awareness could weaken across the secondary school years, even in the lower tracks. As soon as students are placed in a track at the beginning of secondary school, their salient reference group shifts over time from the entire age cohort to only those students in one’s own track 21 . The reality of students’ daily lives and interactions with diverse others in their track could alleviate initial negative beliefs about the image of their track, manifesting in decreasing trajectories for stereotype awareness 22 .
However, considering the current lack of longitudinal research that has explored how students’ awareness of school-track-related stereotypes in different school tracks develops across several years and the contrasting theoretical assumptions about such trajectories, it is difficult to derive clear predictions. Therefore, for the preregistered RQ2 (changes in students’ awareness of school-track-related stereotypes over time in the three different tracks), we did not specify concrete hypotheses and conducted exploratory analyses instead.
Lastly, how is students’ awareness of school-track-related stereotypes related to academic outcomes ( Research Questions 3–5, RQ3-5 )? In the current study, we investigated three types of academic outcomes: Academic achievement in terms of standardized achievement test scores, students’ self-concept of academic aptitude, and school engagement. Below, we outline theoretical considerations and prior research findings on relationships between the awareness of school-track-related stereotypes and academic outcomes, with an emphasis on potential differences between tracks.
The awareness of school-track-related stereotypes may amplify inequalities, as stereotype awareness could be particularly harmful to members of stigmatized groups 23 . Due to the social stigma associated with their track, it is plausible that lower track students’ awareness of negative school-track-related stereotypes negatively affects their school-related development (academic achievement, engagement, self-concept). Potential mechanisms underlying these negative effects among lower track students are, for example, stereotype threat and self-fulfilling prophecies 8 , 24 (but see refs. 3 , 25 ; for research that questioned the relevance of stereotype threat in “real-life” settings). On the other hand, given that higher track students belong to a nonstigmatized (higher status) track, they might not be affected by negative stereotypes that refer to the “typical” higher track student. To conclude, according to what we call the “stereotype awareness as an amplifier of inequality” hypothesis, the awareness of school-track-related stereotypes hampers the thriving of students from the lowest track. Hence, stereotype awareness should be most strongly and negatively related to the academic outcomes of students in the lowest track, thereby increasing the inequalities that are associated with the different school tracks.
Whereas the “stereotype awareness as an amplifier of inequality” hypothesis primarily builds on social psychological and sociological theories, an individual differences perspective supports a competing hypothesis. Such a competing hypothesis could state that group categories and objective status differences associated with different tracks may be less important, and instead, individuals’ subjective perceptions of negative school-track-related stereotypes matter most (see also, e.g., research on effects of subjective socioeconomic status 26 , 27 ). This implies that school-track-related stereotype awareness might not be exclusively maladaptive for students from the lowest track. Rather, stereotype awareness may be negatively related to academic outcomes of individuals from other tracks, too, as long as they believe that such stereotypes exist; an assumption we call the “stereotype awareness as harmful for all” hypothesis.
Regarding links to academic achievement, a third assumption seems plausible. As students are sorted into different tracks on the basis of achievement, two scenarios can be outlined for achievement. Even though stereotype awareness may be negatively related to achievement, it is also reasonable to assume that relatively higher achieving students in a track are particularly likely to devalue their track and generalize this devaluing to how others perceive their track because, for them, the next higher track may also have been an option (from herein labeled the “stereotype awareness as an indicator of missed opportunities” hypothesis). In line with Big Fish Little Pond reasoning 28 such a student could be the “big fish” (a relatively high-achieving student) in a “little pond” (their current track); however, as they did not make it to the “big pond” (next higher track), they devalue their current track and believe that other people in general do so as well.
Prior research on school-track-related stereotype awareness and academic outcomes able to test different theoretical assumptions has been limited. In a sample containing predominantly lower track students (Hauptschule, 72%) but not as many students attending the highest ability track in Germany (Gymnasium, 28%), awareness of school-track-related stereotypes was significantly and negatively correlated with students’ self-concept of academic aptitude, motivation, and academic achievement 7 . However, the cross-sectional design prevented conclusions from being drawn about developmental relationships between stereotype awareness and critical outcomes. In addition, school track was included as a predictor, but relationships between students’ awareness of school-track-related stereotypes and outcome variables were not explored separately for different school tracks 7 .
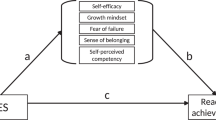
In light of the scarce amount of research on relationships between school-track-related stereotype awareness and academic outcomes, it seemed most appropriate to rely on theoretical considerations (in terms of the above outlined three hypotheses) to guide the current study’s preregistered research questions. Hence, RQ3 (relationships between initial levels of stereotype awareness assessed in Grade 5 right after students transitioned to secondary school and initial levels of academic outcomes), RQ4 (relationships between initial levels of stereotype awareness and developmental trajectories in outcomes variables), and RQ5 (relationships between changes in stereotype awareness and changes in outcome variables), we combined the “stereotype awareness as an amplifier of inequality” hypothesis and the “stereotype awareness as harmful for all” hypothesis. Thus, we hypothesized that (initial levels of and changes in) stereotype awareness should be negatively related to (initial levels of and changes in) all academic outcomes; however, while we assumed that the effects would be present in all investigated tracks (“stereotype awareness as harmful for all” hypothesis), we proposed that effect sizes should be largest for students from the lowest track in line with the “stereotype awareness as an amplifier of inequality” hypothesis. We hypothesized that there would be negative relationships between school-track-related stereotypes and the outcomes, with the potential exception of achievement scores. Whereas we expected to find significant relationships with achievement, we left it open whether these would be negative or, reflecting the “stereotype awareness as an indicator of missed opportunities” hypothesis, positive. Figure 1 provides an overview of all five research questions. Except for RQ1, which was addressed with tests of mean differences in stereotypes between tracks, all research questions were addressed with multigroup growth curve analysis. For RQs 3–5, we controlled for effects of potentially confounding variables (socioeconomic status, gender, migration background). This study’s research questions, hypotheses, and main analyses were preregistered on the Open Science Framework prior to the main analyses (on December 7, 2022, https://osf.io/uwjrm/ ). All analysis codes can also be found on the OSF.
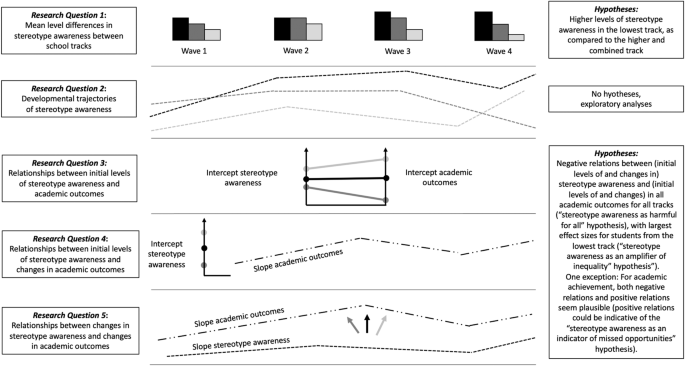
Overview of the five research questions (left side) and hypotheses (right side) addressed in this study.
Bivariate correlations and descriptive information
Bivariate correlations are reported separately for the three tracks in Supplementary Figs. 1 – 3 in the Online Supplement. The student composition in the three tracks showed that students from the lowest track had lower SES backgrounds ( M SES = 43.21, SD = 11.86, measured as the highest socio-economic index of occupational status of the parents, HISEI 29 , which integrates information on income and education and can range from 16 [cleaner] to 90 [judge]) and were more likely to come from a migration background (46.8%) than students in the other two tracks (higher track: M SES = 49.46, SD = 13.44, 16.9% migration background; combined track: M SES = 45.54, SD = 12.11, 4.6% migration background). The gender composition was very similar, with 56%, 55%, and 53% male students in the lowest, combined, and higher tracks, respectively. Descriptive statistics ( M and SD of all variables) and information on missing data can be found in Supplementary Table 1 . We also calculated intraclass correlation coefficients for the stereotype awareness measure separately for each track for the four waves to provide additional information. ICC(1) values ranged from 0.000 to 0.081. Overall, little variance could be attributed to the classroom level, indicating that school-track-related stereotypes are best viewed as an individual-student-level construct and not as a group-level (i.e., classroom-level) phenomenon. Measurement invariance over time and across tracks was tested and generally supported (for details, see the “Method” section).
We deviated from the preregistered main analyses in two significant ways: First, we had preregistered that we could use school belonging as an outcome; however, due to persistent model convergence problems, we decided not to include the results for school belonging in this paper. Second, the analyses were based on a one-factor model for stereotype awareness comprising cognitive, social, and motivational stereotypes and not on separate factors (see the “Method” section for more details on the CFAs for stereotype awareness).
Mean differences in stereotype awareness between tracks (RQ1)
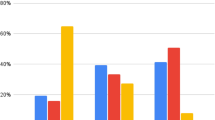
Tests based on model comparisons revealed significant mean differences in latent stereotype awareness factors between tracks for all waves (models with constrained means between tracks had a significantly worse fit than models with unconstrained means, χ 2 (2) ranging from 66.58 to 216.63, all p s < 0.001). Follow-up tests with a Benjamini-Hochberg correction to adjust for multiple testing indicated that students from the lowest track reported significantly higher levels of stereotype awareness in all waves than students from the higher and combined tracks (all p s < 0.001, except for the comparisons between the lowest and combined tracks in Grade 5: p = 0.002, Grade 6: p = 0.003, and Grade 8: p = 0.011). Students from the combined track further reported significantly higher levels of stereotype awareness than students from the higher track in all waves (all p s < 0.001). Figure 2 presents the mean-level differences.
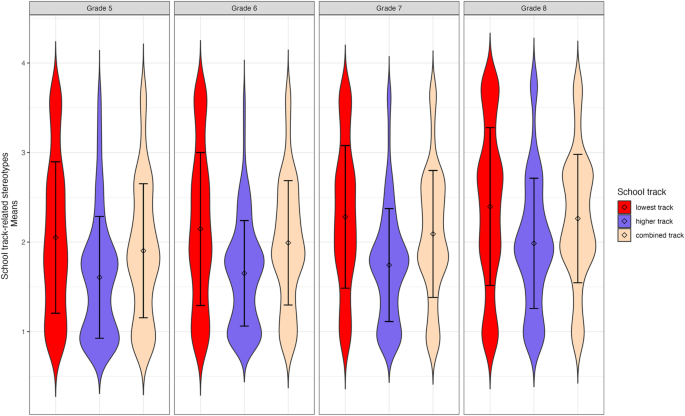
Violin plots displaying mean-level differences (including standard deviations) in school-track-related stereotype awareness between the three secondary school tracks for all four waves.
Developmental trajectories of stereotype awareness (RQ2)
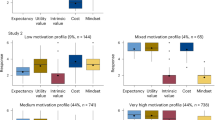
Multigroup univariate growth curve models were set up to investigate developmental trajectories in school-track-related stereotype awareness (see Supplementary Table 5 for details about growth curve model parameters). Stereotype awareness significantly increased in all tracks across the 4-year period (lowest track: b = 0.281; higher track: b = 0.329; combined track: b = 0.288; all p s < 0.001, see Fig. 3 ). To quantify the magnitude of the change, we additionally calculated Glass’s ∆ as an effect size indicator 30 , 31 . Glass’s ∆ amounted to 0.306, 0.447, and 0.371 for the lowest track, the higher track, and the combined track, respectively (all p s < 0.001). These findings indicate that the mean levels of stereotype awareness in all three tracks increased by roughly one third of a standard deviation across the 4 years. Moreover, comparing a model constrained to be equal across tracks with an unconstrained model revealed that the developmental trajectories did not differ significantly between tracks, χ 2 (2) = 0.912, p = 0.634.
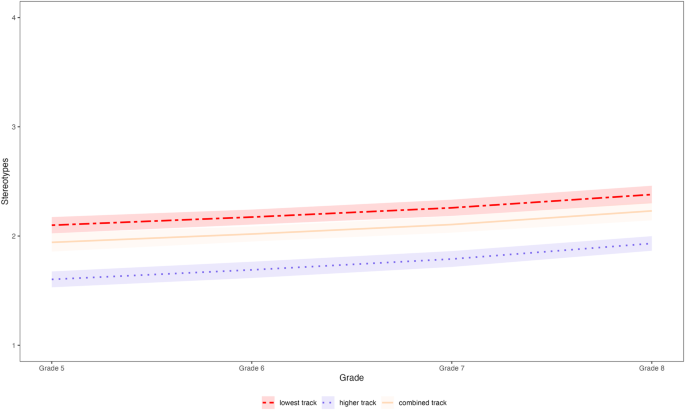
The developmental trajectories (including 95% CIs) in school-track-related stereotype awareness from Grade 5 (first year of secondary school) to Grade 8 in the three tracks.
Relationships between initial levels of and changes in stereotype awareness and academic development (RQ3–RQ5)
We estimated multigroup growth curve models to investigate the relationships between the initial levels of stereotype awareness and academic outcomes (RQ3), relationships between initial levels of stereotype awareness and changes in outcomes (RQ4), and relationships between changes in stereotype awareness and outcomes (RQ5), controlling for SES, gender, and migration background (Fig. 4 ). Separate models were set up for each outcome, and all three research questions were addressed in one model. The results are summarized in Table 1 (school engagement), Table 2 (academic achievement), and Table 3 (self-concept of academic aptitude).
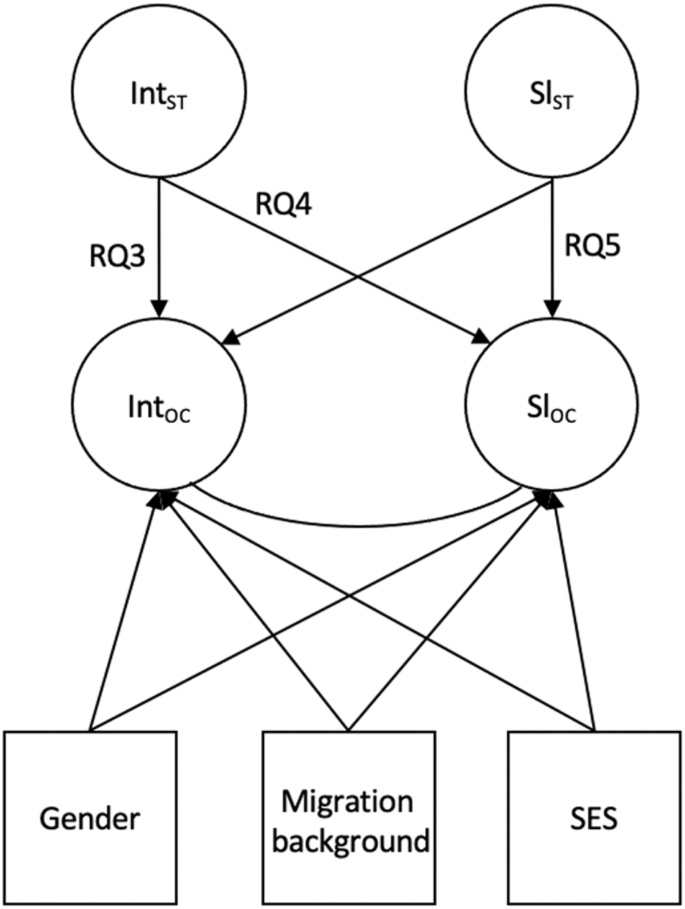
RQ3 addresses relationships between initial levels (intercept) of stereotype awareness and initial levels (intercept) of outcome variables in Grade 5, RQ4 addresses relationships between initial levels (intercept) of stereotype awareness in Grade 5 and changes (slope) in outcome variables, RQ5 addresses relationships between changes (slope) in stereotype awareness and changes (slope) in outcome variables. Int intercept stereotype awareness, Sl ST slope stereotype awareness, Int OC intercept outcome, Sl OC slope outcome. The control variables gender, migration background, and socioeconomic status (SES) are displayed. Correlations between exogenous variables are not shown.
Initial levels of school-track-related stereotype awareness in Grade 5 were significantly and negatively related to initial levels of engagement in the lowest track (b = −0.360, p < 0.001), the higher track (b = −0.594, p < 0.001), and the combined track (b = −0.469, p < 0.001; RQ 3). However, initial levels of stereotypes were not significantly related to students’ development in engagement across the 4 years (RQ 4), and the developmental trajectories of stereotype awareness and engagement were not significantly related in any of the three tracks (RQ 5). Model comparisons revealed that a more parsimonious model in which parameters were constrained to be equal across tracks did not show a significantly worse fit than an unconstrained model, meaning that none of the effects differed significantly between tracks, χ 2 (8) = 6.259, p = 0.618.
Achievement
Initial levels of school-track-related stereotype awareness in Grade 5 were significantly and positively related to initial levels of achievement in the lowest track (b = 0.647, p < 0.001), whereas the relationships were not statistically significant in the two other tracks (RQ3, see Table 2 ). Initial levels of stereotypes were further significantly and negatively related to developmental trajectories in achievement, when initial levels of achievement were controlled for, in the combined track (b = −0.314, p = 0.046; RQ4). Developmental trajectories in stereotype awareness were not significantly related to developmental trajectories in achievement in any of the tracks (RQ5). Results from model comparisons indicated that the constrained model had a significantly worse fit than the unconstrained one, χ 2 (8) = 50.983, p < 0.001, revealing significant differences between tracks. Follow-up tests with the Benjamini-Hochberg correction indicated that for RQ3, the effect in the lowest track differed significantly from effects in the higher track and the combined track (both p s < 0.001). The combined and higher tracks did not differ significantly ( p = 0.128). For RQ4, no differences between tracks were found (lowest track vs. higher track: p = 0.738; lowest track vs. combined track: p = 0.738; combined track vs. higher track: p = 0.884). For RQ5, follow-up tests showed significant differences between the higher track and the combined track ( p = 0.038). The effect in the lowest track did not differ significantly from the effects in the higher track ( p = 0.673) or the combined track ( p = 0.244).
Self-concept of academic aptitude
Initial levels of school-track-related stereotype awareness in Grade 5 were significantly and negatively related to initial self-concept levels in the higher track (b = −0.251, p = 0.001; RQ3). None of the relationships between initial stereotype awareness and changes in self-concept were statistically significant. Changes in stereotype awareness were not significantly related to changes in self-concept. There were no statistically significant differences in the effects between tracks, χ 2 (8) = 12.427, p = 0.133.
Secondary school tracking generates inequalities in opportunities and has pervasive consequences for individuals’ educational trajectories and life paths 32 . Despite the widespread use of tracking in Germany and other countries and efforts to identify psychological barriers that further widen school-track-related inequalities in education 11 , stereotypes have received surprisingly little attention to date 3 , 7 , 9 , 33 . The present 4-year longitudinal study on students’ awareness of negative school-track-related stereotypes therefore advances current knowledge in important ways.
We found significant mean-level differences in school-track-related stereotype awareness between all three tracks for all waves in the expected direction, with students from the lowest track consistently reporting higher levels of stereotype awareness than those from the higher and combined tracks. These findings present an important replication and extension of prior work 7 , documenting significant differences between students from the lowest track (Hauptschule, also included in our data) and students from Gymnasium, the highest ability track in Germany. Our finding also add to differentiation-polarization theory 9 , 10 , 12 , 15 by suggesting that the polarization component can be applied to students’ awareness of negative school-track-related stereotypes, with the most negative perceptions in the lowest track.
Further, negative school-track-related stereotypes increased in all tracks. From a developmental perspective, secondary school tracking coincides with the onset of adolescence, a period in which students’ cognitive capacities to understand the social implications of their academic placement grow 11 . A potential explanation for these ascending trajectories that could apply to students from all three of the tracks we investigated stresses the fact that school-track-related stereotype awareness referring to one’s own school track likely develops in reference to other tracks. Over time and as the end of compulsory schooling nears, restricted opportunities (e.g., for students’ further education and career) linked to their current track as compared with the next higher track(s) may become more apparent to students. Importantly, this phenomenon should also apply to many students from the relatively highest track in our study, who still tend to have fewer opportunities than students from Gymnasium, the highest academic track in Germany, which was not represented in our data.
Next, we investigated links between stereotype awareness and academic outcomes. To summarize the main findings, we did not find support for any of our hypotheses longitudinally, as changes in students’ stereotype awareness were not related to changes in their academic achievement, self-concept of academic aptitude, or engagement.
It was shown that stereotype awareness in Grade 5 significantly predicted a more maladaptive development of achievement over the course of 4 years in the combined track, but the effects did not differ significantly between tracks. Cross-sectionally, we obtained some limited evidence for the “stereotype awareness as harmful for all” hypothesis but only for student engagement. With respect to self-concept of academic aptitude, we found a significant negative cross-sectional relationship to stereotype awareness for students in the higher track; however, the effects for self-concept were not significantly different across the three tracks. Moreover, a positive relationship between stereotype awareness in Grade 5 and achievement was obtained for the lower track, and this effect differed significantly from the effects in the higher track (a nonsignificant small negative relationship) and combined track (a nonsignificant small positive relationship). The cross-sectional effect for academic achievement in the lower track was in line with the “stereotype awareness as an indicator of missed opportunities” hypothesis. None of the obtained effects confirmed the assumptions outlined in the “stereotype awareness as an amplifier of educational inequalities” hypothesis.
Jointly, our findings are consistent with the view that stereotype awareness goes along with lower levels of some aspects of academic functioning, as indicated by negative cross-sectional relationships with engagement in all tracks and a negative cross-sectional relationship with self-concept of academic aptitude in the higher track. The findings for engagement revealed that students who started secondary school with higher levels of negative stereotype awareness were less engaged and reported enjoying school less. Although the effect for self-concept did not differ across tracks, it suggests that for higher track students, the awareness of negative stereotypes about their track was ingrained into their self-perceptions to a higher degree at the beginning of secondary school. Students in the higher track in our data (“Realschule”) were probably more likely to better understand their position and the implications of their track placement, including the awareness that they were not assigned to the highest ability track (i.e., Gymnasium). This understanding could strengthen the link between stereotype awareness and self-concept found in Grade 5. The explanation resembles the mechanisms outlined in the integration paradox. The integration paradox describes the phenomenon that relatively more highly educated immigrants turn away from the host society instead of becoming more oriented toward it 34 .
The cross-sectional positive relationship with academic achievement for students from the lowest track is in contrast with the negative cross-sectional relationships for engagement and self-concept and casts new light on relationships between school-track-related stereotype awareness and achievement 7 , 9 . We interpret this effect as providing support for the “stereotype awareness as an indicator of missed opportunities” hypothesis. Specifically, the awareness of negative school-track-related stereotypes may serve as an indicator of missed opportunities such that higher achieving students, for whom the next higher track could also have been an option due to their relatively higher achievement levels, report higher levels of awareness of stereotypes that refer to their current track. A reason for why significant effects were restricted to the lowest track students could be that for relatively high-achieving students from the lowest track, positive and self-worth-protecting comparison processes that students from other tracks could use (“I might not have made it to the next higher track, but at least I am not in the next lower one!”) do not work. Therefore, missed educational opportunities may be perceived as particularly drastic 35 and could go hand in hand with negative school-track-related stereotypes. A potential explanation for why the same pattern was not obtained for students from the combined track, which represents the relatively lower track in the federal state of Saxony, could be that the combined track offers different educational opportunities and types of leaving exams. Therefore, students from this track may feel less “stuck,” and thus, they are less prone to suffer from the mechanisms outlined in the “stereotype awareness as an indicator of missed opportunities” hypothesis. Of course, at this point, our explanations are purely speculative and should be tested empirically in future work.
Finally, stereotype awareness in Grade 5 significantly predicted a more maladaptive development of achievement over the course of 4 years in the combined track, a finding that may indicate that initial stereotype awareness signals resignation and disappointment with one’s track placement that could feed into poorer performance over time. Nonetheless, the sizes of the effects in the three tracks were very similar, and, despite the differentiated pattern of significant and nonsignificant results, the effects did not differ significantly between tracks. Even though we did not obtain any significant relationships between developmental trajectories in stereotype awareness and developmental trajectories in achievement or any of the other academic outcomes, it is worth mentioning that the negative effect for achievement in the higher track just failed to reach statistical significance and differed significantly from the effect in the combined track.
Our study has implications for theory and the understanding of how school-track-related stereotype awareness operates in real life. In short, it is complex and differs from what would be expected on the basis of related research in the laboratory (e.g., on stereotype threat 8 ). These differences are likely due to the multifaceted experiences that students have in complex social environments, dynamic shifts in environments and reference groups, students’ subjective interpretations thereof, along with (variations in) developmental processes during adolescence. Hence, we propose that research and future theory development regarding school-track-related stereotype awareness cannot afford to be blind to “the context,” adolescents’ respective meaning making, and dynamic shifts in their perceptions. Further, a developmental (longitudinal) perspective is crucial for disentangling concurrent and longitudinal associations. Relatedly, a more extensive formulation of a theory of school-track-related stereotype awareness in real-life contexts is also informed by what we did not find. Specifically, clarity on the potential implications of stereotype awareness in terms of consistent longitudinal relationships with the investigated outcomes could not be achieved. Nonetheless, given the, to the best of our knowledge, lack of longitudinal research on stereotype awareness spanning several years of adolescents’ school careers, the present study’s findings significantly contribute to the existing body of research. What we now need are context-sensitive developmental studies that can capture stereotype awareness on different time scales. For example, longer-term longitudinal assessments could be combined with intensive longitudinal assessments at critical time points (e.g., the transition to secondary school) to shed light on dynamic changes in stereotype awareness, implications for academic outcomes, and interactions with changes in comparison processes and reference group effects.
Our work has implications for practice and policy too. In the present study, students in the lowest track reported the highest levels of stereotype awareness in all waves. Hence, we see a need to counteract school-track-related stereotypes in daily interactions with students and in the media. Still, efforts to curb stereotypes should not be constrained to the lowest track, as relatively higher tracks may also be negatively affected (e.g., Realschule in our data). Moreover, we caution that it is not only about improving the image of tracks but also about improving the actual life realities and chances for students in these tracks. In Germany, secondary school tracking occurs early, track-related upward mobility is still very limited, and the permeability of the education system likely interacts with family background characteristics 35 . In addition to streaming students at a later point in their educational careers, coaching and other types of interventions 36 —especially prior to and at the transition to secondary school and with a focus on socioeconomically disadvantaged students and their parents—could provide a remedy. Students who missed the next higher track could be another relevant target group for interventions, as in our study, relatively higher achieving students in the lowest track reported higher levels of negative school-track-related stereotype awareness right after starting secondary school.
Several limitations and directions for future research should be noted. First, in our study, stereotype awareness was assessed solely in reference to a student’s own track. Future research could gain insights into social comparison processes by investigating how these stereotype awareness ratings differ from students’ awareness of stereotypes about other school tracks. Relatedly, it would be interesting to explore the extent to which the perception that a much stronger stigma is associated with one’s track than with other tracks (i.e., larger stereotype awareness gaps) drives relationships between stereotypes and outcome variables. Second, the awareness of negative stereotypes was assessed and analyzed as a single, general construct, which ensured comparability of the stereotype construct across tracks. At the same time, however, this measurement approach falls short of capturing the existence of potentially different types of stereotypes for students in different tracks. For example, stereotypes about students in higher tracks may be that they are nerdy, boring, socially awkward, elitist, “know-it-alls” or teacher’s pets 7 . Future research should thus include a larger number of more diverse school-track-related stereotypes. Nonetheless, we believe that the content of the stereotype measure we employed was still suitable for our study. Although in our study, the mean levels of stereotype awareness were highest in the lowest track (Hauptschule), significant relationships between stereotype awareness and the outcome variables were in several instances obtained for (or even restricted to) students from Realschule, the higher track. This latter finding indicates that the negative stereotypes we assessed are relevant to higher track students. Third, substantially distinct stereotypes likely exist for the highest ability track (Gymnasium), but this track was not represented in our data.
Fourth, the present study focused on outcomes that reflect academically relevant features in three different domains (academic achievement: actual performance; self-concept of academic aptitude: motivational self-beliefs; engagement: affective school involvement) and that are arguably proximal to academically relevant stereotypes (e.g., “stupid,” “not interested in school”). On the other hand, this focus necessarily excluded other critical outcomes. Future studies on school-track-related stereotype awareness should expand the scope of the present investigation to explore relationships between school-track-related stereotype awareness and a range of outcomes in the socioemotional (e.g., well-being), social (e.g., social networks), emotional (e.g., academic emotions), and behavioral (e.g., disruptive behavior, school drop-out) domains.
Fifth, several items from the academic self-concept scale measure asked students to evaluate their abilities in comparison with others. However, the measure did not specify who these “others” were. Future studies should enhance clarity by indicating and systematically contrasting different reference groups (e.g., elementary school classmates, new secondary school classmates, friends).
Sixth, lastly, limitations that refer to the composition of the sample can be identified and used to inform future studies. Even though we were able to take advantage of a large and rich data set with four measurement points that spanned students’ lower secondary school careers, it remains a drawback that students from the highest track in Germany (Gymnasium) were not included due to the focus of the TRAIN study on specific school types. In addition, differences in secondary school systems across the German federal states need to be kept in mind when interpreting the findings. Specifically, the comparison between Hauptschule (lowest track) and Realschule (higher track) from the federal state of Baden-Württemberg is unequivocal, whereas caution is warranted when comparing these two tracks with Mittelschule in Saxony.
To conclude, stratified school systems likely create stereotypes about students attending different tracks 7 . Our study showed that students from the lowest track consistently reported higher levels of negative school-track-related stereotype awareness than students attending the higher and combined tracks. However, the findings on links between the awareness of school-track-related stereotypes and academic outcomes did not indicate that school-track-related stereotype awareness amplifies educational inequalities. As an initial longer term longitudinal investigation of school-track-related stereotype awareness exploring patterns of effects in different school tracks, our study notably adds to the literature and contributes to a more differentiated understanding of students’ awareness of the stigma associated with their school track.
We used previously collected data from a large-scale longitudinal German study (TRAIN) with four measurement points, which is hosted by the Hector Research Institute of Education Sciences and Psychology at the University of Tübingen in Germany ( https://uni-tuebingen.de/en/43704 ). The TRAIN study was based on a multistage sampling design, in which school-type-specific subpopulations of interest (from the three tracks Hauptschule, Realschule, and Mittelschule) were drawn disproportionately to the actual population shares. The TRAIN study relied on stratified cluster samples, which were drawn separately for both participating federal states (Baden-Württemberg, Saxony). First, a random sample of schools (cluster) was determined for both federal states. From each school, fifth-grade classes were then randomly selected, and all students in these classes were asked to participate in the TRAIN study. This sampling strategy was employed due to the TRAIN study’s focus on the three school tracks of Hauptschule, Realschule, and Mittelschule and allowed for detailed analyses of students and schools from these three tracks. However, the sample is therefore not representative of the population (Rose et al. 37 ). The sampling procedure used by the TRAIN study resulted in a target sample of 22 schools for Mittelschule, 25 schools for Realschule, and 60 schools for Hauptschule, all of which were contacted and invited to participate. Out of the schools that were invited, one school from the Realschule track and one school from the Hauptschule track did not participate. The student response rates for the survey assessments were 83–89%, 90–93%, and 74–80% in the Hauptschule, Realschule, and Mittelschule tracks, respectively 37 .
The sample analyzed in this study contained data from 3880 secondary school students enrolled in 136 classes from two German federal states (Baden-Württemberg, 66%, and Saxony, 34%), for whom data from at least one measurement point was available. Across all measurement points, 45.2% of the students identified as female, and they were, on average, 14.20 years old at the fourth measurement point ( SD = 0.65). A total of 43% of the students attended the academically least demanding track (lowest track, Hauptschule), 23% attended the higher track (Realschule), and 34% attended the combined track (Mittelschule). The students from Hauptschule and Realschule in our sample were from the German federal state of Baden-Württemberg. Most German states, including Baden-Württemberg, track students in the threefold school system consisting of Gymnasium (the highest track), Realschule (the next higher track), and Hauptschule (the lowest track). Hauptschule, the school track with the lowest academic demands, is mainly vocationally oriented. The Realschule curriculum is focused on general education but also lays the foundation for future vocational careers. In comparison with Hauptschule, Realschule provides students with much better opportunities for acquiring higher educational qualifications. Lastly, Gymnasium represents the academically most demanding track, and successfully completing Gymnasium entitles students to study at university 2 , 38 . In addition, the sample included students from the German federal state of Saxony attending a combined track, Mittelschule. Students from Mittelschule could acquire a Hauptschule diploma or a Realschule diploma. It should further be noted that, like Baden-Württemberg, the secondary school system in Saxony also includes Gymnasium. However, there are no Realschule or Hauptschule tracks in Saxony. Hence, for our study, comparisons between students from Hauptschule (referred to as the lowest track) and Realschule (referred to as the higher track) from Baden-Württemberg are straightforward; however, any comparisons between these two tracks and Mittelschule (referred to as the combined track) should take into consideration the fact that the students from Mittelschule came from another German federal state (Saxony) with a slightly different secondary school system. All variables used in this study were assessed four times, when students were on average 11 (Grade 5), 12 (Grade 6), 13 (Grade 7), and 14 (Grade 8) years of age. All assessments took place some weeks after the start of the respective school year. The study was approved by the state authorities of Baden-Württemberg and Saxony, who, at this time, were responsible for approving studies like this one. Parental consent was required for study participation.
Achievement was measured with standardized tests. Stereotypes, self-concept of academic aptitude, and school engagement were assessed via student reports on a 4-point scale ranging from 1 ( completely disagree ) to 4 ( completely agree ). Students’ migration background was measured with student reports, whereas family socioeconomic status was captured with parent reports. Gender was assessed with student and teacher ratings combined into one variable (e.g., in cases in which students did not report their gender, information from teacher reports was used).
Awareness of school-track-related stereotypes
We used a measure of negative school-track-related stereotypes referring to a student’s own secondary school track 7 . The items were introduced with the following phrase: “What do you think? How do other people in general think about students attending [your school track]? Other people think that typical students from [your school track] are ….” Students were asked to rate the “typical student” from their school track on nine adjectives referring to negative stereotypes with cognitive (“stupid,” “unimaginative,” “dumb”), motivational (“unmotivated in class,” “not interested in school,” “lazy”), and social content (“rude,” “cheeky,” “brazen”). Reliability coefficients (Cronbach’s alpha) for the awareness of school-track-related stereotypes measure for the four waves were .921, .929, .918, and .942 for students from the lowest track; .916, .896, .914, and .935 for students from the higher track; and .932, .923, .929, and .935 for students from the combined track, respectively.
Academic achievement
Two indicators of academic achievement were used, namely, standardized achievement test scores in mathematics and German. The tests included standard content from the federal states’ mathematics curricula (e.g., arithmetic rules, linear equations, and angels) and German language curricula (i.e., reading comprehension). Open ended, closed ended, and multiple-choice response formats were used (for more detailed descriptions, see refs. 39 , 40 ). Item and person parameters for students’ mathematics and German achievement have previously been estimated with longitudinal, multidimensional, two-parameter item response theory models 37 , and we relied on weighted likelihood estimators (WLEs) of students’ mathematics and German achievement test scores 41 , that is, one indicator for each subject and wave. To capture school achievement more broadly, we built an average of students’ mathematics and German test scores for the analyses of this study.
Students’ self-concept of academic aptitude was measured with four reverse-coded items (“Frequently, I’m convinced that I won’t be able to solve a task even before I get started”; “I frequently think that I’m not as smart as others are”; “I’d like to be as intelligent as others are”; “Compared with others, I’m not as talented”) 42 . Reliability coefficients (Cronbach’s alpha) for the four waves were .668, .714, .727, and .748 for students from the lowest track; .717, .712, .773, and .776 for students from the higher track; and .735, .779, .772, and .776 for students from the combined track, respectively.
School engagement
We used three items to map school engagement. The items were based on the BIJU study (“I enjoy working on my tasks at school”; “In the morning, I look forward to a day at school to learn something new”; “School is a place I enjoy being at”) 43 . Reliability coefficients (Cronbach’s alpha) for the four waves were .746, .727, .729, and .740 for students from the lowest track; .775, .824, .769, and .765 for students from the higher track; and .782, .756, .769, and .770 for students from the combined track, respectively.
The control variables gender (0 = female , 1 = male ), migration background (0 = no migration background , 1 = migration background ), and socioeconomic background (SES; highest socio-economic index of occupational status of the parents, HISEI 30 ) were considered. Specifically, these control variables were included in all analyses in which effects on academic outcomes were estimated (i.e., the analyses for RQ3–RQ5).
Measurement models and measurement invariance testing
We tested the factor structure of all multiple-item scales with confirmatory factor analyses (CFAs) conducted in M plus Version 8.6 44 , including longitudinal measurement invariance testing and testing for invariance between school tracks. For growth curve models (main analyses), at least scalar invariance (equal item intercepts and factor loadings) is needed 45 . For stereotypes, we furthermore compared models with different numbers of stereotype awareness factors. Originally, scholars distinguished between students’ awareness of cognitive, social, and motivational school-track-related stereotypes 7 , and for our study, we tested for whether these stereotypes represent (a) three distinct facets, (b) one overarching construct, or (c) two constructs by comparing three-factor, two-factor, and one-factor CFA models. We assessed the goodness of fit of all models using the comparative fit index (CFI), the Tucker-Lewis Index (TLI), and the root mean square error of approximation (RMSEA). Typical cut-off scores that are considered to reflect excellent and adequate fit to the data, respectively, were considered: (a) CFI and TLI > .95 and >.90, (b) RMSEA < 0.05 and <0.08 46 .
The evaluation of longitudinal and intergroup invariance assumptions was based on respective recommendations from the methodological literature 45 , 47 , 48 . Hence, we considered drops in the CFI or TLI > 0.01 and increases in the RMSEA > 0.015 as indicative of meaningful changes in model fit, which make assumptions of measurement invariance untenable. When relying on latent factors, several problems can occur, and it is not possible to anticipate all of them. For instance, there may be single items that show a substantially low(er) loading on the latent factors than others, and model fits might not be in line with traditional recommendations 46 . When this was the case, we carefully checked both statistical indicators (e.g., lower loading, worse model fit) and content-related indicators (e.g., a specific item might not “represent” the respective construct as well as other items do) and adapted our models to achieve adequate fit. Accordingly, for school engagement, we decided to use the three (positively worded) items from the original seven-item scale that best captured students’ broader affective school engagement based on the respective CFA (and additionally conducted exploratory factor analysis) results and conceptual considerations. Moreover, for the school engagement scale, residuals from the same items were allowed to correlate across time.
For stereotype awareness, the CFA findings showed that a two-factor model with a factor for cognitive stereotype awareness and a factor combining social and motivation stereotype awareness fit the data slightly better than the next best one-factor solution (two-factor model: CFI = .954, TLI = .949, RMSEA = 0.031; one-factor model: CFI = .942, TLI = .938, RMSEA = 0.034). The three-factor model did not converge. In addition, it should be noted that we only included the negatively worded items because including both positively worded (reverse-coded) and negatively worded items resulted in a very poor model fit and convergence problems (introducing a method factor did not solve the problem). Conceptually, as our aim was to measure the awareness of negative stereotypes, focusing on the negatively worded items seems appropriate, and we therefore excluded the three positively worded statements for cognitive, social, and motivational stereotype awareness, respectively (e.g., “smart,” “polite”). There was a deviation from our preregistered analysis plan for the main analyses (see the “Statistical analyses” section for the main analyses). On the basis of the CFA results, we had preregistered that we would model cognitive and social/motivational stereotype awareness separately. However, despite the slightly superior fit of the respective two-factor solution in comparison with a one-factor solution, a closer inspection revealed that the two stereotype awareness factors were strongly correlated. Whereas the manifest correlations ranged from .78 to .83 (which was slightly lower than those previously reported 7 , the latent correlations ranged from .90 to .97. We therefore decided to rely on one overall stereotype awareness factor in our (latent) main analyses. Tables reporting detailed CFA and measurement invariance testing results for all constructs can be found in the Supplementary Tables 2 – 4 . To summarize, both longitudinal invariance and invariance between tracks could generally be established, and the main analyses (see Statistical analyses) were based on the respective models.
Statistical analyses
All analyses were performed with M plus Version 8.6 44 using the robust maximum likelihood estimator (MLR), which is robust to non-normal data. To deal with missing data, we employed full information maximum likelihood estimation (FIML 49 ). All multiple-item scales were modeled as latent variables. Achievement was modeled as a manifest indicator for which mathematics and German test scores were combined.
For RQ1, we tested for mean level differences in stereotype awareness between the three tracks separately for each wave. Specifically, we tested for mean differences in latent stereotype awareness factors by constraining them to equality between groups and then comparing the constrained model with a model with unconstrained stereotypes. This comparison was done jointly, and then we compared individual groups as a follow-up test (including the adjustment of p -values with the Benjamini-Hochberg correction).
Next, we set up multigroup latent growth curve models. To investigate developmental trajectories in school-track-related stereotype awareness (RQ2), we estimated univariate growth models. To allow greater flexibility for the shape of the curve, we estimated latent basis growth models in which only the first and the last growth parameters were fixed, whereas all other parameters were freely estimated. Thereby, no linearity was imposed on the models. However, the growth parameters were set to be equal between the groups (tracks) so that we could meaningfully compare the slopes between them.
For RQ3–RQ5, we estimated multigroup growth curve models to examine relationships between initial levels (intercepts) of stereotype awareness and initial levels (intercepts) of all outcomes (RQ3), relationships between initial levels (intercepts) of stereotype awareness and changes (slopes) in all outcomes (RQ4), as well as relationships between changes (slopes) in school-track-related stereotype awareness and changes (slopes) in all outcome variables (RQ5). We created separate models for each outcome due to model complexity. All models included time-invariant covariates (SES, gender, migration background). We evaluated differences in parameters between the groups by using a nested-models χ 2 approach with follow-up comparisons to isolate the source of the overall differences. To account for the hierarchical data structure, with students nested in classes, the analyses were conducted with cluster-robust standard errors. All significance testing was performed at the 0.05 level, and we relied on two-tailed tests. Benjamini-Hochberg corrections 50 were used to adjust for multiple tests. We applied the adjustment for each research question separately (but across the three different outcomes) and adjusted all p -values that were relevant for our research questions 51 , 52 .
Data availability
The data are currently not publicly available due to data privacy/ethical restrictions. Data used for this submission may be made available upon request to collaborators of the TRAIN study research team ( https://uni-tuebingen.de/en/faculties/faculty-of-economics-and-social-sciences/subjects/department-of-social-sciences/education-sciences-and-psychology/research/aktuelle-studien/train/ ).
Code availability
All analysis code is available on the OSF (anonymized link: https://osf.io/uwjrm/?view_only=ef06f871bcb34447929c8ac34fe277f1 ).
Becker, M., Lüdtke, O., Trautwein, U., Köller, O. & Baumert, J. The differential effects of school tracking on psychometric intelligence: do academic-track schools make students smarter? J. Educ. Psychol. 104 , 682–699 (2012).
Article Google Scholar
Maaz, K., Trautwein, U., Lüdtke, O. & Baumert, J. Educational transitions and differential learning environments: how explicit between‐school tracking contributes to social inequality in educational outcomes. Child Dev. Perspect. 2 , 99–106 (2008).
Chu, J., Loyalka, P., Li, G., Gao, L. & Song, Y. Stereotype threat and educational tracking: a field experiment in Chinese vocational high schools. Socius 4 , 1–11 (2018).
Batruch, A., Geven, S., Kessenich, E. & van de Werfhorst, H. G. Are tracking recommendations biased? a review of teachers’ role in the creation of inequalities in tracking decisions. Teach. Teach. Educ. 123 , 1–18 (2022).
Google Scholar
Durante, F. & Fiske, S. T. How social-class stereotypes maintain inequality. Curr. Opin. Psychol. 18 , 43–48 (2017).
Article PubMed PubMed Central Google Scholar
Domina, T., Penner, A. & Penner, E. Categorical inequality: schools as sorting machines. Annu. Rev. Sociol. 43 , 311–330 (2017).
Knigge, M. & Hannover, B. Collective school-type identity: predicting students’ motivation beyond academic self-concept. Int. J. Psychol. 46 , 191–205 (2011).
Article PubMed Google Scholar
Steele, C. A threat in the air: how stereotypes shape intellectual identity and performance. Am. Psychol. 52 , 613–629 (1997).
Article CAS PubMed Google Scholar
Spruyt, B., Van Droogenbroeck, F. & Kavadias, D. Educational tracking and sense of futility: a matter of stigma consciousness? Oxf. Rev. Educ. 41 , 747–765 (2015).
Hargreaves, D. H. Social Relations in a Secondary School . (Routledge, London, 1967).
Legette, K. A. Social-cognitive perspective of the consequences of curricular tracking on youth outcomes. Educ. Psychol. Rev. 32 , 885–900 (2020).
Van Houtte, M. Lower-track students’ sense of academic futility: selection or effect? J. Sociol. 52 , 874–889 (2016).
Negru-Subtirica, O., Pop, E. I. & Crocetti, E. Developmental trajectories and reciprocal associations between career adaptability and vocational identity: a three-wave longitudinal study with adolescents. J. Vocat. Behav. 88 , 131–142 (2015).
Major, B. & O’Brien, L. T. The social psychology of stigma. Annu. Rev. Psychol. 56 , 393–421 (2005).
Van Houtte, M. School type and academic culture: evidence for the differentiation–polarization theory. J. Curric. Stud. 38 , 273–292 (2006).
Abraham, J. Testing Hargreaves’ and Lacey’s differentiation-polarisation theory in a setted comprehensive. Br. J. Sociol. 40 , 46–81 (1989).
Garcia-Marques, L., Santos, A. S. C. & Mackie, D. M. Stereotypes: static abstractions or dynamic knowledge structures? J. Pers. Soc. Psychol. 91 , 814–831 (2006).
Cromley, J. G. et al. Changes in race and sex stereotype threat among diverse STEM students: relation to grades and retention in the majors. Contemp. Educ. Psychol. 38 , 247–258 (2013).
Totonchi, D. A., Perez, T., Lee, Y., Robinson, K. A. & Linnenbrink-Garcia, L. The role of stereotype threat in ethnic minority students’ declining science motivation: a four-year longitudinal study of achievement and persistence in STEM. Contemp. Educ. Psychol. 67 , 102015 (2021).
Rivenbark, J. G. et al. Perceived social status and mental health among young adolescents: evidence from census data to cellphones. Dev. Psychol. 55 , 574–585 (2019).
Chmielewski, A. K., Dumont, H. & Trautwein, U. Tracking effects depend on tracking type: an international comparison of students’ mathematics self-concept. Am. Educ. Res. J. 50 , 925–957 (2013).
Konovalova, E. & Le Mens, G. An information sampling explanation for the in-group heterogeneity effect. Psychological Rev. 127 , 47–73 (2020).
Swencionis, J. K. & Fiske, S. T. Stereotypes and Relative Social Status in Social Comparisons. In Social Comparison, Judgment, and Behavior (eds Suls, J., Collins, R. L. & Wheeler, L.) 251–279 (Oxford University Press, Oxford, 2020).
Madon, S. et al. The accumulation of stereotype-based self-fulfilling prophecies. J. Pers. Soc. Psychol. 115 , 825–844 (2018).
Cullen, M. J., Hardison, C. M. & Sackett, P. R. Using SAT-grade and ability-job performance relationships to test predictions derived from stereotype threat theory. J. Appl. Psychol. 89 , 220–230 (2004).
Rivenbark, J. G. et al. Adolescents’ perceptions of family social status correlate with health and life chances: a twin-difference longitudinal cohort study. Proc. Natl Acad. Sci. USA 117 , 23323–23328 (2020).
Article CAS PubMed PubMed Central Google Scholar
Tan, J. J. X., Kraus, M. W., Carpenter, N. C. & Adler, N. E. The association between objective and subjective socioeconomic status and subjective well-being: a meta-analytic review. Psychol. Bull. 146 , 970–1020 (2020).
Marsh, H. W. & Parker, J. W. Determinants of student self-concept: Is it better to be a relatively large fish in a small pond even if you don’t learn to swim as well? J. Pers. Soc. Psychol. 47 , 213–231 (1984).
Ganzeboom, H. B., De Graaf, P. M. & Treiman, D. J. A standard international socio-economic index of occupational status. Soc. Sci. Res. 21 , 1–56 (1992).
Miyamoto, A., Murayama, K. & Lechner, C. M. The developmental trajectory of intrinsic reading motivation: measurement invariance, group variations, and implications for reading proficiency. Contemp. Educ. Psychol. 63 , 101921 (2020).
Morris, S. B. & DeShon, R. P. Combining effect size estimates in meta-analysis with repeated measures and independent-groups designs. Psychol. Methods 7 , 105–125 (2002).
Knigge, A., Maas, I., Stienstra, K., De Zeeuw, E. L. & Boomsma, D. I. Delayed tracking and inequality of opportunity: gene-environment interactions in educational attainment. NPJ Sci. Learn 7 , 1–13 (2022).
Legette, K. School tracking and youth self-perceptions: implications for academic and racial identity. Child. Dev. 89 , 1311–1327 (2018).
Verkuyten, M. The integration paradox: empiric evidence from the Netherlands. Am. Behav. Scientist 60 , 583–596 (2016).
Forster, A. G. Caught by surprise: the adaptation of parental expectations after unexpected ability track placement. Res. Soc. Stratif. Mobil. 76 , 100630 (2021).
Easterbrook, M. J. & Hadden, I. R. Tackling educational inequalities with social psychology: Identities, contexts, and interventions. SIPR 15 , 180–236 (2021).
Rose, N. et al. Durchführung und Methodische Grundlagen der TRAIN-Studie. In Tradition und Innovation: Entwicklungsverläufe an Haupt- und Realschulen in Baden-Württemberg und Mittelschulen in Sachsen - Abschlussbericht für die Länder Baden-Württemberg und Sachsen (eds Jonkmann K., Rose N. & Trautwein U.) 77–102 (2013).
Frenzel, A. C., Goetz, T., Pekrun, R. & Watt, H. M. Development of mathematics interest in adolescence: Influences of gender, family, and school context. J. Adolesc. Res. 20 , 507–537 (2010).
Aldrup, K., Klusmann, U., Lüdtke, O., Göllner, R. & Trautwein, U. Social support and classroom management are related to secondary students’ general school adjustment: a multilevel structural equation model using student and teacher ratings. J. Educ. Psychol. 110 , 1066–1083 (2018).
Dumont, H., Trautwein, U., Nagy, G. & Nagengast, B. Quality of parental homework involvement: predictors and reciprocal relations with academic functioning in the reading domain. J. Educ. Psychol. 106 , 144–161 (2014).
Warm, T. A. Weighted likelihood estimation of ability in item response theory. Psychometrika 54 , 427–450 (1989).
Schwanzer, A. D., Trautwein, U., Lüdtke, O. & Sydow, H. Entwicklung eines Instruments zur Erfassung des Selbstkonzepts junger Erwachsener. Diagnostica 51 , 183–194 (2005).
Baumert, J. et al. Bildungsverläufe und psychosoziale Entwicklung im Jugendalter (BIJU): Dokumentation Unpublished Manuscript. (1997).
Muthén, L. K. & Muthén, B. O. Mplus User’s Guide . Eighth Edition . https://www.statmodel.com/download/usersguide/MplusUserGuideVer_8.pdf (1998–2017).
Little, T. D., Card, N. A., Slegers, D. W. & Ledford, E. C. Representing Contextual Effects in Multiple-Group MACS Models. In Modeling Contextual Effects in Longitudinal Studies (eds Little, T. D., Boivard J. A. & Card, N. A.) 121–147 (Routledge, Oxfordshire, 2007).
Hu, L.-T. & Bentler, P. M. Cutoff criteria for fit indexes in covariance structure analysis: conventional criteria versus new alternatives. Struct. Equ. Model 6 , 1–55 (1999).
Chen, F. F. Sensitivity of goodness of fit indexes to lack of measurement invariance. Struct. Equ. Model. 14 , 464–504 (2007).
Cheung, G. W. & Rensvold, R. B. Evaluating goodness-of-fit indexes for testing measurement invariance. Struct. Equ. Model 9 , 233–255 (2002).
Enders, C. K. Applied Missing Data Analysis (Guilford Press, New York, 2010).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc., Ser. B. Stat. Methodol. 57 , 289–300 (1995).
Lakens, D. Sample size justification. Collabra Psychol. 8 , 33267 (2022).
Matsunaga, M. Familywise error in multiple comparisons: disentangling a knot through a critique of O’Keefe’s arguments against alpha adjustment. Commun. Methods Meas. 1 , 243–265 (2007).
Download references
Acknowledgements
This study’s research questions, hypotheses, and main analyses were preregistered on the Open Science Framework prior to the main analyses (on December 7, 2022, https://osf.io/uwjrm/ ). All analysis codes can also be found on the OSF. The TRAIN study was funded by the Ministry of Education and Cultural Affairs Baden Württemberg, the Robert-Bosch foundation and the Hertie-foundation. L.B. and K.M. are supported by Jacobs Foundation Research Fellowships, and K.M. is supported by an Alexander von Humboldt Professorship endowed by the German Federal Ministry of Education and Research. We acknowledge support from the Open Access Publication Fund of the University of Tübingen.
Open Access funding enabled and organized by Projekt DEAL.
Author information
Authors and affiliations.
University of Tübingen, Hector Research Institute of Education Sciences and Psychology, Tübingen, Germany
Lisa Bardach, Claudia Neuendorf, Kou Murayama, Thorsten Fahrbach, Benjamin Nagengast & Ulrich Trautwein
University of Potsdam, Education Department, Potsdam, Germany
Claudia Neuendorf
Humboldt University Berlin, Department of Rehabilitation Sciences, Berlin, Germany
Michel Knigge
Korea University, Department of Education and the Brain & Motivation Research Institute (bMRI), Seoul, Republic of Korea
Benjamin Nagengast
You can also search for this author in PubMed Google Scholar
Contributions
Conceptualization: L.B., C.N., K.M., B.N., U.T.; Methodology: C.N., L.B., B.N., K.M.; Validation: C.N., T.F.; Formal analysis: C.N.; Writing—Original draft: L.B., C.N.; Writing—Review and editing: All authors; Visualization: C.N., T.F.; Resources: M.K.; Funding acquisition: U.T.
Corresponding author
Correspondence to Lisa Bardach .
Ethics declarations
Competing interests.
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary figures and tables, rights and permissions.
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/ .
Reprints and permissions
About this article
Cite this article.
Bardach, L., Neuendorf, C., Murayama, K. et al. Does students’ awareness of school-track-related stereotypes exacerbate inequalities in education?. npj Sci. Learn. 8 , 59 (2023). https://doi.org/10.1038/s41539-023-00203-9
Download citation
Received : 24 December 2022
Accepted : 15 November 2023
Published : 14 December 2023
DOI : https://doi.org/10.1038/s41539-023-00203-9
Share this article
Anyone you share the following link with will be able to read this content:
Sorry, a shareable link is not currently available for this article.
Provided by the Springer Nature SharedIt content-sharing initiative
This article is cited by
Using social and behavioral science to address achievement inequality.
- Eddie Brummelman
- Nienke van Atteveldt
- Jellie Sierksma
npj Science of Learning (2024)
Grade 12 students’ perceptions of educational tracks in Flanders
- Margo Vandenbroeck
- Jonas Dockx
- Rianne Janssen
Social Psychology of Education (2024)
Quick links
- Explore articles by subject
- Guide to authors
- Editorial policies
Sign up for the Nature Briefing newsletter — what matters in science, free to your inbox daily.
- Academic Tracking: Long-term Effects on College and Career Outcomes

- All Rights Reserved
- Woods, Kenton Bentley (Author)
- Hanish, Laura (Thesis advisor)
- DeLay, Dawn (Committee member)
- Martin, Carol (Committee member)
- Jager, Justin (Committee member)
- Arizona State University (Publisher)
- Educational Administration
- Educational philosophy
- academic tracking
- post-secondary education
- Partial requirement for: Ph.D., Arizona State University, 2023
- Field of study: Family and Human Development
View full metadata

Machine-readable links
- OAI Dublin Core
Quick actions
Academia.edu no longer supports Internet Explorer.
To browse Academia.edu and the wider internet faster and more securely, please take a few seconds to upgrade your browser .
Enter the email address you signed up with and we'll email you a reset link.
- We're Hiring!
- Help Center

FACTORS THAT AFFECTS GRADE 10 STUDENTS IN CHOOSING ACADEMIC TRACK IN SENIOR HIGH SCHOOL

Factors Affecting Grade 10 Students from Maligaya High School in choosing Academic Track for Senior High School for the School Year 2019-2020 In partial fulfillment for the subject Practical Research 1 To be presented for the faculty members of ACCESS COMPUTER AND TECHNICAL COLLEGES - Lagro Campus By: (List the name of each member alphabetically)
Related Papers
Rahser Mahinay
Daniel Fil Divino
With the changes that are needed to be faced by our country in terms of educational curriculum, the researchers have made a move to pursue this study. In our study, we concluded four (4) major factors which was the basis of this study, Parental Influence, Aptitude, Interests and Environmental Factors. This study aims to find out the significant differences between the career choice factors and the gender of our respondents. The research was conducted at the University of the Immaculate Conception and its respondents were selected Grade 10 students, ranging from 20-23 per section. It was performed using the descriptive survey method, thus, the researchers formulated a questionnaire based on the four (4) different indicators, with six (6) statements each. The questionnaires were distributed in 8 sections, with 172 respondents all in all which was verified through the Slovin’s formula. The researchers then encoded the data to be able to get the mean scores, as well as, the p-value or the significant difference. It was then formulated by the SPSS, and obtained a p-value of 0.144. Therefore, it was implied that there was a significant difference between the career choices of grade 10 students with their gender. The proponents’ decision was to accept the alternative hypothesis and reject the null hypothesis. There are diverse and several factors which can also affect the career choice of an individual. For the improvement of further studies, the researchers highly recommend that there should be other factors that will be looked upon since career choice is essential in one’s future way of life.
justin arenas
Jefferson Oraño
This study aims to determine the factors that affect the senior high school track preferences of the Grade 9 students of Don Bosco Technology Center of academic year 2014-2014. This study utilizes descriptive method of research to determine the factors. It would see if dependent variables relating to personality, family/relatives, interests and job opportunities were significant factors influencing the track preferences of the respondents. The descriptive research used quantitative methods to assess the feedback from the respondents
IRA International Journal of Education and Multidisciplinary Studies
jerald moneva
There are many influences that affect the preferences of grade 10 students in choosing a track to proceed to senior high school. Likewise, this study aims to identify influence of preference of a Senior High School track that is commonly encountered by the Grade 10 students in terms of Gender, Socio-Economic Status, Average academic grades, nature of parent’s occupation; and, strand and the level of influence of the respondent to be associated with preferences in choosing a track in senior high school in terms of family influence-decision; peer influence; financial condition; and employability. The research tool was a survey questionnaire. The questionnaire is composed of respondent’s profile and 10 statements to be rated. The factors fairly influence preferences of the senior high school. In terms of gender, male students consider their socio-economic status and their parent’s occupation as factors in choosing their track in Senior High School while female students consider thei...
Rudy Daling
The study highlighted the transition rate of Grade10-Junior High School (JHS) completers to Grade 11-Senior High School (SHS) enrolment, and students’ track preferences. The study utilized quanti-quali approach. Quantitative data were collected and analyze from school’s records/forms. Qualitative data were based from direct responses of the respondents. It employed descriptive-evaluative type of research design and applied purposive sampling to 41 students to obtain descriptions out from the results of evaluation. The high percentage distribution of “Balik-aral” students, classified as parent-students, contributed an increase of Grade 11 enrolment. The career goals of SHS program, College and Business/Commerce, encouraged the community to patronize to study the track offered by the school. Students’ mastery level did not suffice the passing standard in education that the SHS track offered by the school was not relevant to the preferences of students. Thus, the school may assess or evaluate school’s program and curriculum instruction suited to learners’ learning style to prepare them according to their level of preferences. KEYWORDS: Students’ Preferences, SHSTrack, Curriculum, Transition Rate
Joshua O Japitan
This study aims to determine the factors that affect the senior high school track preferences of the Grade 9 students of Don Bosco Technology Center of academic year 2014-2014. This study utilizes descriptive method of research to determine the factors. It would see if dependent variables relating to personality, family/relatives, interests and job opportunities were significant factors influencing the track preferences of the respondents. The descriptive research used quantitative methods to assess the feedback from the respondents. Scale/questionnaire is given to the respondents to conduct the study personally and is collected after to gather all the results. Most of the literature gathered talks about the factors that affect career preferences/choices, namely personality, family, interests and job opportunities, which would specialized in senior high school of the K-12 curriculum.
Psychology and Education: A Multidisciplinary Journal
Psychology and Education
The study aimed to determine the factors affecting senior high school track preferences of Grade 10 students in the district of Morong. The respondents of the study were 495 students which is 50 percent of the total population of grade 10 students in the said school. The study revealed that the respondents were mostly females belonging to family with monthly income below ₱10,000. Several are fourth child in the family whose fathers were college undergraduates and whose mothers were high school graduates. The perceived extent of the factors influencing the Grade 10 students in their senior high school track preferences with respect to personal, family, peer, and school was found to be Much; however, with respect to community the extent is found to be Moderate. The null hypothesis was rejected for the significant difference on the perception of the students on the extent of the factors affecting their track preferences in terms of their sex, sibling position, monthly family income and fathers' educational attainment. Meanwhile, the null hypothesis was accepted for the significant difference on the perception of the students on the extent of the factors affecting their track preferences in terms of their mothers' educational attainment.
kim venus dotado
Career selection is one of the most important choices that students will make in determining their future plans.The Philippines is one of the remaining country in ASIA with a 10 years of school in secondary level of education. This short period of time makes difficult for every Filipinos to become competitive with other Nations who at least have 12 years of basic education. Holland (2010) individuals are attracted to a given career by their particular personalities and numerous vauiables that constitute their backgrounds. First of all career choice is an expression of or an extension of
Loading Preview
Sorry, preview is currently unavailable. You can download the paper by clicking the button above.
RELATED PAPERS
International journal of Academic Research in Education and Review
Oscar Vallejo
Janice Gentoral
Sindani Philip
Sapienza: International Journal of Interdisciplinary Studies
Maurice dence Bacaling
Cognizance Journal of Multidisciplinary Studies
Lorrein Jashmin M. Maglalang
Eumi Cuntapay
Neil Cumigad
British Journal of Education Society Behavioural Science
mohammed sulemana
Marianne Roxas-Villanueva
Jenneriche Corpus
Lambert Academic Publishing
Justice Agyei Ampofo
Nordin Abd Razak
Lander Amparo
International Journal of Educational Research Review
Yasin BAYRAKTAR
Journal of Vocational Education Studies
Muhammad Sayuti
Sylvester Onyeka
AL-ISHLAH: Jurnal Pendidikan
eza aldonna
Indonesian Journal of Educational Research and Technology
Rezel Dagamac
Halimatus Sakdiah
Fritz Gerald Martin
Shar-In A. Sadjail , Darwina J. Sansawi , Mona-Allea L. Matolo , Psychology and Education
Cyreanth Trujulan
RELATED TOPICS
- We're Hiring!
- Help Center
- Find new research papers in:
- Health Sciences
- Earth Sciences
- Cognitive Science
- Mathematics
- Computer Science
- Academia ©2024
The Research Whisperer
Just like the thesis whisperer – but with more money, building your track record.

She can be found on Twitter at @deborahbrian , where she talks higher education policy, research strategy, Australian politics, social justice, and cats. Mostly cats.
A version of this article first appeared in Funding Insight on December 14, 2017 and is reproduced with kind permission of Research Professional. For more articles like this, visit www.researchprofessional.com .

As the year begins, many of you will be planning your research for the coming year and identifying funding schemes to target. Some will have received the outcomes of last year’s grant applications and will either be breathing a sigh of relief or girding their loins for the next attempt.
This can be a difficult time, both professionally and emotionally, for early career researchers in particular (see Tseen Khoo’s recent post on academic disappointment ).
This is especially so for those in fields where there is an expectation that salaries will be sourced from grant and fellowship funds.
In this era of short-term contracts and reduced security of employment, there has never been more pressure on early career researchers to establish a research track record.
Couple this with declining grant success rates across the board and increasing competition and the situation can become quite daunting. Those who are not successful in becoming one of the 1 in 10 researchers awarded a major grant or fellowship can easily become disheartened.
Some tell me the major funding bodies just don’t care about their field, are biased against their particular methodology, or that it is all a lottery anyway. None of this is true, of course, but – more importantly – it isn’t helpful.
So, what can you do if you are an early career researcher struggling to break into the big leagues of research funding?
Here are five tips for you to help build your track record:
Start small
Major grants tend to be very heavily dependent on track record, so even if you are a talented researcher with a brilliant idea, you may not get funded without a strong track record of past research.
One way to build that track record is to apply to smaller, less competitive funding schemes, such as those offered by charitable foundations or internal grants and fellowships administered by your institution.
While these are generally of a lower value and less prestigious, they give you the opportunity to fund smaller research projects, such as pilot studies, and demonstrate your capacity to successfully manage a research project to produce outcomes. Both of these things will stand you in good stead in the next funding round to which you apply.
Think outside the box
In addition to pursuing small grant opportunities, consider ways you can make your own funded research opportunities. Establish and maintain strong networks with relevant industry groups, government departments, and others, and identify areas where you might collaborate on contract or consultancy research. Talk to your institution’s research donations staff or directly to relevant charitable groups to see if there are donors willing to support your research.
Again, you may need to start small, but even small projects will help you build your project management skills and track record, and you never know where they might lead. Some researchers build whole careers based on the support of charitable foundations, particularly in medical research. Others develop productive industry partnerships that support large teams over many years.
As well as networking with other interested parties and end users, be sure to take opportunities to attend seminars and conferences, and get to know your fellow researchers. If you lack the funding to get out to many of these events, get online and participate in the growing academic and disciplinary communities there (this may have the side benefit of developing your own capacity for science communication, outreach, and engagement).
Find like-minded people, and those with complementary skill sets, both in your own cohort and those slightly more senior. Look to your PhD advisors, senior colleagues, and mentors to help you find your way onto active research teams. Not only are teams more likely to be funded in most contexts, they can take on more ambitious research and produce a larger number of outputs, thus contributing to the track record of all co-investigators.
Joining existing teams can be a fast track to funding success and advancement, but collaborative networks are also an important part of your own academic capital.
Tend your own garden
While gaining research funding is important for enabling your research, and, for better or worse, has become a research performance metric in its own right, there are other ways to grow your track record. The most obvious is, of course, publication.
When early career researchers learn they have missed out on that grant or fellowship and ask “What now?”, my answer is invariably: publish. Spend some time writing up existing research and developing it for publication in appropriate journals (you will notice I recommend ‘appropriate’ journals and not necessarily ‘leading journals’, but that’s a whole other story…).
So, write, publish, apply for prizes, attend conferences, build networks, hone your skills, curate your online profile, and carry on tending your garden, building your research capability and finding ways to express that in ways that matter to your communities.
Finally, try to identify ways to move your research forward without funding, whether that is reviewing the literature, analysing existing data sets, or running a pilot experiment on borrowed lab supplies.
Sell your story
My last tip is in some ways the most important: No matter how strong your track record, it won’t help you if you hide your light under a bushel.
I talk to so many early career researchers who undersell themselves because their track record is not yet as developed as it might be in ten years’ time. The challenge early in your career is to identify potential directions, and distinguish yourself from others.
How do you do that if your track record is a little on the ‘thin’ side? You look to quality, impact, and relative performance. Is your track record of publications strong for someone of your career stage in your field? Get specific. How many of your publications are first author or sole author publications? You may not have many citations yet, but are some of them in top journals? Has your work been featured in the media (including social media), has it generated a lot of clicks, views, downloads, likes, shares? Has your research generated impact in your field, in policy, or in the community?
Don’t forget context! Does your track record represent an ideal preparation to take on this project ? If you designed the project, it really should. Tell your story in a way that shows how the stepping stones of your track record led you directly to this point, and how this project will move you on to the next step.
While you’re waiting for your track record to grow, remember that showcasing a developing research career trajectory in this way can be an ideal strategy when applying for early career fellowships and prizes.
Three to remember
So, as we return to our desks after a well-earned holiday break, remember these tips:
- Be realistic when choosing the funding schemes to which you will apply, and balance the time you spend writing grant applications with time spent writing papers and ‘tending your garden’.
- Spend some time looking for alternative funding opportunities , and building networks with other researchers and end users.
- Make time to think about how you represent your track record , exploring the bibliometric tools at your disposal for analysing and benchmarking your publication record, and following up citations and other measures of reach and impact.
Above all, remember that there are as many pathways to a successful research career as there are successful researchers. Which path will you choose?
Share this:
One comment.
Reblogged this on Digital learning PD Dr Ann Lawless .
Leave a comment Cancel reply
This site uses Akismet to reduce spam. Learn how your comment data is processed .
- Already have a WordPress.com account? Log in now.
- Subscribe Subscribed
- Copy shortlink
- Report this content
- View post in Reader
- Manage subscriptions
- Collapse this bar
Establishing an Academic Track Record
Cite this chapter.

- Lorrie Blair 2
Part of the book series: Teaching Writing ((WRIT))
8044 Accesses
Not long ago, new graduates were hired on the basis of their promise. Today, they need proven track records that include conference presentations and peer-reviewed publications. They cannot wait for graduation to start building their Curriculum Vitae (CV), as competition for grants, post-doctorate positions, and jobs require that students are already active in their fields.
This is a preview of subscription content, log in via an institution to check access.
Access this chapter
Subscribe and save.
- Get 10 units per month
- Download Article/Chapter or eBook
- 1 Unit = 1 Article or 1 Chapter
- Cancel anytime
- Available as PDF
- Read on any device
- Instant download
- Own it forever
Tax calculation will be finalised at checkout
Purchases are for personal use only
Institutional subscriptions
Unable to display preview. Download preview PDF.
Author information
Authors and affiliations.
Concordia University, Canada
Lorrie Blair
You can also search for this author in PubMed Google Scholar
Rights and permissions
Reprints and permissions
Copyright information
© 2016 Sense Publishers
About this chapter
Blair, L. (2016). Establishing an Academic Track Record. In: Writing a Graduate Thesis or Dissertation. Teaching Writing. SensePublishers, Rotterdam. https://doi.org/10.1007/978-94-6300-426-8_9
Download citation
DOI : https://doi.org/10.1007/978-94-6300-426-8_9
Publisher Name : SensePublishers, Rotterdam
Online ISBN : 978-94-6300-426-8
eBook Packages : Education Education (R0)
Share this chapter
Anyone you share the following link with will be able to read this content:
Sorry, a shareable link is not currently available for this article.
Provided by the Springer Nature SharedIt content-sharing initiative
- Publish with us
Policies and ethics
- Find a journal
- Track your research
Longitudinal Academic Tracks

Our Longitudinal Academic Tracks allow students to explore various areas of interest in conjunction with the four-year medical school curriculum. All tracks are faculty-run, scholarly experiences for medical students interested in developing attitudes and skills for self-directed, lifelong learning and career development.
The goals of the Longitudinal Academic Tracks are to promote intellectual curiosity, appreciation of scholarly inquiry, inter-professional collaboration, and cura personalis .
Longitudinal Academic Tracks currently offered in the School of Medicine:
- Diversity, Equity, & Inclusion in Medicine Track
- Environmental Health and Medicine Track
- Health Justice Scholar Track
- Healthcare Leadership Track
- Literature and Medicine Track
- Medical Education Research Scholar Track
- Population Health Scholar Track
- Primary Care Leadership Track
- Spirituality in Medicine Track
- Bioethics Academic Track
Interested students may apply during first year of medical school. Students who successfully complete a track by meeting all track requirements and in good academic standing, graduate from the medical school with distinction.
Students may apply to up to two academic tracks. The track application becomes available to students in late Fall of their first year.
The Longitudinal Academic Tracks are extracurricular opportunities that enhance the core curriculum and allow students to build upon a specific interest in medicine over the course of the four-year undergraduate medical education curriculum. In effort to ensure prioritization of core curricular elements and support workload balance, students must be in good academic standing (as outlined below) to participate in a Track.
Any student below passing in two longitudinal courses, or below passing in one longitudinal course and below 70% in two or more modules, will NOT be eligible to participate in an extracurricular track. Longitudinal courses that are less than 10% complete at the end of Block 2 are not considered in the determination of good academic standing for track participation eligibility.
- Toggle Accessibility Statement
- Skip to Main Content

Academic Track
| /Research/Career Advocacy/Culminating Activity i.e. | |
| / /Research/Career Advocacy/ | |
| / /Career Advocacy/Culminating Activity | |
| /Research/Career Advocacy/Culminating Activity | |
| /Research/Career Advocacy/Culminating Activity |
| *Select from HUMSS Strand Subjects 1 to 4. **Select from HUMSS Strand Subjects 5 to 8. ***Schools must present/offer a range of subjects from which students can choose. |
Academic Research Track (ART)
The Academic Research Track (ART) was a program for first, second and third year medical students who considered a research experience as part of their education or career.
- Past Projects
- Education Directory
ART seminars introduce research concepts, useful in planning a mentored summer or 'year out' research project. Taught in small lunchtime seminars (light lunch included) occur throughout the medical school year. Modules consisting of two to three seminars each cover such topics as mentoring and being mentored, formulating a research question, measurements, data gathering and presentation, obtaining research funding, and writing for publication. The goal is to help students think about science as they plan for a mentored research experience, which is typically taken after the second or third years of medical school.
The ART program administrator is Jene Dupra .
The faculty contact is Candace Gildner .
Current ART Trainees
Antoinette Nguyen
Sumeetha Swaminathan
Past ART Trainees
Baqir Kedwai Project: Determination of the Natural History of Aortic Dissection Tissue Mechanics Using Non-invasive Elastography Mentor: Doran Mix, M.D.
Daniel Lehane Project: Patient-Reported Outcomes for Ruptured Abdominal Aortic Aneurysm Mentor: Karina Newhall, M.D., M.S.
Tori Valachovic Project: Assessing Reproductive Health Care and Perspectives amongst Birthing People with Epilepsy Mentor: Sarah Betstadt, M.D., M.P.H.
Marissa LoCastro Project: Engagement in Advanced Care Planning Among Older Patients with Acute Myeloid Leukemia and Myelodysplastic Syndrome Mentor: Dr. Melissa (Kah Poh) Loh
Caroline Maretz Project: Infectious Keratitis: Epidemiology, Bacterial Virulence and Antimicrobial Resistance Mentor: Dr. Rachel Wozniak
Ching-Wei Pan Project: Molecular Mechanism of GATA1 Mutation Causing Transient Abnormal Myelopoiesis in Down Syndrome Mentor: Dr. Laurie Steiner
Mariah Erlick Project: Assessing Value in the Surgical Episode of Care: A Conceptual Model and Implementation Framework Aligning Patient and Provider Values in Gastrointestinal Oncologic Surgery Mentor: Dr. Larissa Temple
Eleanor Pope Project: Longitudinal Real World Changes in Skin Microbial Ecology in AD Patients Mentor: Dr. Lisa Beck
Kevin Vo Project: Targeting Beta-II Spectrin to Improve CAR-T Cell Therapy Mentor: Dr. Minsoo Kim
Shireen Saxena Project Title: "Sex and Race Differences in Outcomes for Patients with Primary Prevention Implantation of Cardioverter Defibrillators" Mentor: Ilan Goldenberg, M.D.
Racquel Whyte Project Title: "Investigating the Difference in Platelet Phenotypes in Patients with Acute Ischemic Stroke and Large Vessel Occlusion versus Non-Stroke Controls" Mentor: Matthew Bender, M.D.
Fatima Bawany Project Title: “Understanding the Relationship Between Skin Commensal Bacterial and S. aureus Burden in Patients with Atopic Dermatitis” Mentor: Lisa Beck, M.D.
Alejandra Rodriguez Project Title: “The Effects of Growth Hormone and IGF-1 on Retinal Nerve Fiber Layers Following Compression Injury” Mentors: G. Edward Vates, M.D., Ph.D. and David Paul, M.D., M.S.
Victor Wang Project Title: “Examining the Relationship of Cigarette Smoke and the Development of Proliferative Vitreoretinopathy” Mentor: Ajay Kuriyan, M.D., M.S.
John Wilson Project Title: “Envelope-Following Response (EFR) as a Modality for Assessing Cochlear Synaptopathy in the Budgerigar” Mentor: Kenneth Henry, Ph.D.
Timothy Campbell Project Title: Look Like an Expert: Gaze Training in Novices Enhances Rate of Skill Acquisition in a VR Simulated Robotic Suture Task Mentor: Ahmed Ghazi, M.D., M.Sc.
Samuel Tomlinson Project Title: Role of interictal spike activity in planning the resection margins for epilepsy surgery Mentor: Eric Marsh, M.D., Ph.D. at the Children’s Hospital of Philadelphia
Kwanza Warren Project Title: Early Post-surgical Temozolomide Therapy in Patients with High-Grade Gliomas Admitted to Acute Rehabilitation: A Feasibility Study Mentor: Kevin A. Walter, M.D.
Justin Williams Project Title: The Association Between Health Literacy, Health Outcomes, and Medication Adherence in Patients with Multiple Sclerosis Mentors: Jessica Robb, M.D. and Andrew Goodman, M.D.
Asad Arastu Project Title: Association of Financial Toxicity (FT) with Depression, Anxiety, and Quality of Life (QoL) in Older Patients and Caregivers with Advanced Cancer Mentor: Supriya Mohile, M.D., M.S.
Shravani Gangidi Project Title: Heart Rate Variability Among ECG and PPG Signals as predictor of Atrial Fibrillation Recurrence After Electrical Cardioversion Mentor: Jean-Philippe Couderc, Ph.D., M.B.A.
Rachel Park Project Title: Identification of Novel Small Molecules that Prevent Radiation-Induced Capsular Contracture Mentor: Richard Phipps, Ph.D.
Michelle Shankar Project Title: Depression among Urban Teens with Asthma: A Focus for Providers Mentor: Jill Halterman, M.D., M.P.H.
Earlier Years
Jennifer Andreozzi Project Title: Geriatric Assessment to Improve Outcomes of Elderly Cancer Patients Mentor:Supriya Mohile, M.D., M.S.
Josef Bartels Project Title: The Context, Structure, and Function of Silence in Doctor-Patient Communication Mentor: Ronald Epstein, M.D.
Jon Black Project Title: How We See It: Life through the Eyes of a Pediatric Cancer Patient Mentor: Nancy Chin, Ph.D., M.P.H.
Jarrod Bogue Project Title: Investigation of the fundamental biochemistry and conformational properties of a specific riboswitch from Neisseria gonorrhoeae Mentor: Joseph Wedekind, Ph.D.
Zachary Borus Project Title: The Impact of Implementing a 'Collaborative Problem Solving' Approach to Care on an Inpatient Child Psychiatry Unit Mentor: David Garrison, M.D.
Matthew Brown Project Title: Molecular, Cellular, and Tissue Abnormalities Leading to Impaired Fracture Healing in C57BL/6J Marine Model of Type II Diabetes Mentor: Regis O'Keefe, M.D., Ph.D.
Pedro Calderon-Artero Project Title: The Effects of Potent Inflammation Resolving Lipid Mediators on Platelet Function in Healthy and Diabetic Blood Mentor: Robert Block, M.D., M.P.H., F.A.C.P.
Amanda Carpenter Project Title: Aldo-keto reductase (AKR1C3) Role in Basal & Squamous Cell Skin Cancer Diabetic Blood Mentor: Alice Pentland, M.D.
Jennifer Cialone Project Title: A Phase II, Randomized, Placebo Controlled Trial of the Safety & Tolerability of Mycophenolate (CellCept) in Children with Juvenile Neuronal Ceroid Lipofuscinosis (JNCL) Mentor: Jonathan Mink, M.D., Ph.D.
Ian DeAndrea-Lazarus Project Title: Early Life Lead Exposure and Executive Functions Using the Stroop Day-Night Task Mentor: Todd Jusko, Ph.D.
Thomas Fugate Project Title: Detecting LVH by ECG in African Americans, Jackson Heart Study Mentor: Thomas Pearson, M.D., M.P.H., Ph.D.
Robert Fulton Project Title: Resolvins as Novel Regulators of Alveolar Lung Epithelial Cell Inflammatory Responses Mentor: Patricia Sime, M.D.
Michael Geary Project Title: Modulation of the prostanoid receptor EP4 to reduce scarring during flexor tendon healing Mentor: Regis O'Keefe, Ph.D.
Trevor Hansen Project Title: Thy1 Expression as a Marker and Therapeutic Target for Scar Formation in Capsular Contracture following Reconstruction Mammoplasty Mentor: Richard Phipps, Ph.D.
Christopher Hogan Project Title: Mechanisms for Protecting Human Lung Fibroblasts from Cigarette Smoke-Induced Cell Death: Implications for COPD Mentor: Patricia Sime, M.D.
Jing-Li Huang Project Title: Autophagy in Pancreatic Ductal Adenocarcinoma Mentor: Aram Hezel, M.D.
Brian Jenssen Project Title: Dissemination of Best Practices to Promote Smoke Free Homes Mentor: Jonathan Klein, M.D., M.P.H.
Evan Katzel Project Title: The impact of Smad3 loss of function on TGF-beta signaling and the resultant scarring and adhesion formation during tendon healing Mentor: Regis O'Keefe, M.D., Ph.D.
Angel Kirkham Project Title: Belize: A Case Study of a Community Mental Health Program in a Developing Nation Mentor: Nancy Chin, Ph.D., M.P.H.
Ryan Koehler Project Title: Development and Validation of the Arthroscopic Surgery Skill Evaluation Tool (ASSET) Mentor: Gregg Nicandri, M.D.
Sirish Kondabolu Project Title: Development of siRNA to Prevent Scar Formation in Tendon Repair Mentor: Regis O'Keefe, M.D., Ph.D.
Ajay Kuriyan Project Title: Inhibition of corneal fibroblast differentiation to scar forming cells Mentor: Richard Phipps, Ph.D.
Chase Kwon Project Title: A Systems Biology Approach to Identify Determinants of Staphylococcus Mentor: Lisa Beck, M.D.
Erika Levy Project Title: Treatment of Anxiety and Obsessive Compulsive Behaviors in Tourette Syndrome Mentor: Jonathan Mink, M.D., Ph.D.
Judy Liu Project Title: Mutation specific risk stratification of Long QT Syndrome type 3 via measure of evolutionary conservation of amino acids expressed by the SCN5A gene Mentor: Arthur Moss, M.D.
Jinno (Tony) Magno Project Title: Increased Sensitivity of Poor-Prognosis Human MLL-Rearranged Acute Lymphoblastic Leukemia to Proteasome Inhibition (Bortezomib) in Vitro Mentor: Jessica Shand, M.D.
Kelly Makino Project Title: Advance Care Planning in Early Dementia Study Mentor: Anton Porsteinsson, M.D.
Andrew Marky Project Title: Activated Protein C is Neuroprotective and Mediates New Blood Vessel Formation and Neurogenesis after Controlled Cortical Impact Mentor: Berislav Zlokovic, M.D., Ph.D.
Alexi Matousek Project Title: The Molecular Pathogenesis of Barrett's Esophagus Mentor: Jeffrey Peters, M.D.
Ron Menorca Project Title: Determining EPO's role in accelerating functional recovery of meme in a mouse chronic neuropathy model Mentor: John Elfar, M.D.
Doran Mix Project Title: Computer Assisted Surgery Tool for Hemodialysis Fistulas (CAST-HDF) Mentor: Ankur Chandra, M.D.
Aimee Morris Project Title: Clinical Characteristics, Musical Variables, and Pathophysiology of Focal Embouchure Dystonia: A Disabling Disorder of Musicians Mentor: Jonathan Mink, M.D., Ph.D.
Gregory Ouellet Project Title: Effect of Sex Hormones on Clinical Course of Patients with Congenital Long QT Syndrome Mentor: Arthur Moss, M.D.
Allison Panzer Project Title: Breastfeeding and Health Outcomes in Low and Very Low Birth Weight Infants at 9 Months of Age. Mentor: Cynthia Howard, M.D., M.P.H.
David Paul Project Title: Using DTI to measure changes in occipital lobe white matter after decompression of the optic chiasm Mentors: Brad Mahon, M.D. and Edward Vates, M.D., Ph.D.
Salvador Peña Project Title: Do Astrocytes Contribute to the Exceptionally High Computational Power of the Human Brain? Mentor: Maiken Nedergaard, M.D., DMSc
Clifford Pierre Project Title: The Role of Altered Cerebrospinal Fluid Dynamics after Brain Surgery Mentor: Maiken Nedergaard, M.D., DMSc
Daniel Platt Project Title: Phenotypic Variability in and Treatment of Episodic Ataxia Type 1 Mentor: Robert Griggs, M.D.
Michael Prucha Project Title: Healthcare Worker's Role in Tobacco Control in the Dominican Republic Mentor: Deborah Ossip, Ph.D.
Yaoli Pu Project Title: P75 Neurotrophin Receptor Signaling and Trafficking Mentor: Nina Schor, M.D., Ph.D.
Izad-Yar Daniel Rasheed Project Title: The Kindling Model of Epilepsy and Wavelet-Based Seizure Detection Methods Mentor: David Pinto
Jeffrey Reed Project Title: The Effects of Erythropoietin on Healthy Bone Mentor: John Elfar, M.D.
Kyle Rodenbach Project Title: Crystatin-C-based renal reserve in children with history of hemolytic uremic syndrome-associated acute kidney injury Mentor: George Schwartz, M.D.
Lauren Roussel Project Title: Evaluating Upper Extremity Function Following Mastectomy in Reconstructed and Non-Reconstructed Women with Breast Cancer
Elizabeth Saionz Project Title: Post-concussion progesterone decline in female athletes Mentor: Jeffrey Bazarian, M.D., M.P.H.
Glenn Schneider Project Title: Inner Ear Gene Therapy Steps to Treat Age-Related Hearing Loss Mentor: Robert Frisina, Ph.D.
Jordan Schramm Project Title: Maternal iron-deficiency and its impacts on auditory temporal processing and myelination in the offspring – translation of animal models into the human system Mentor: Anne Luebke, Ph.D.
Erika Snow Project Title: The Role of E-cigarettes as a Barrier to Smoking Cessation Mentor: Scott McIntosh, Ph.D.
Melissa Squires Project Title: Analysis of Inpatient Data on Patients with Intellectual and Developmental Disabilities at Strong Memorial Hospital Mentor: Stephen Sulkes, M.D.
Laurel Stevens Project Title: Role of Plexin B1 receptor in Melanoma Tumor Progression Mentor: Glynis Scott, M.D.
Brandon Stein Project Title: The Relationship Between Subconcussive Head Trauma and Markers of Brain Injury Mentor: Jeff Bazarian, M.D., M.P.H.
Leigh Sundem Project Title: Erythropoietin for Compression Neuropathy: Preclinical Efficacy and Cellular Site of Action Mentor: John Elfar, M.D.
Otto Thomas Project Title: Characteristics, frequency, and implications of extranodal non-Hodgkin's lymphoma Mentor: Louis Constine, M.D.
Heidi Thompson Project Title: Research into Health Care in the Deaf Population Mentor: Steven Barnett, M.D.
Lindsay Wahl Project Title: One Protein, Multiple Functions: The Role of Tissue Transglutaminase in Pulmonary Fibrosis Mentor: Patricia Sime, M.D.
Joanna Walker Project Title: Central Venous Catheter Thrombosis in Children Mentor: Norma Lerner, M.D.
Andrew Walters Project Title: Identification and Deployment of a Novel Drug to Enhance Cardiac Ischemic Preconditioning Mentor: Paul Brookes, Ph.D.
Adam Weis Project Title: Corneal Wound Healing and Ocular Optics After Descemet's Stripping Endothelial Keratoplasty (DSEK) Mentor: Holly Hindman, M.D.
Sam Weisenthal Project Title: Predicting Acute Kidney Injury in Re-Hospitalized Patients Mentor: Martin Zand, M.D., Ph.D.
Pamela White Project Title: Evaluating the Impact of eRecord on Nursing Practice and Patient Outcomes Mentors: Gail Ingersoll, Ed.D., R.N., FAAN, FNAP
Paul Youn Project Title: Effect of Total Body Irradiation and Pre-Transplant Pulmonary Status on Non-Relapse Mortality after Hematopoietic Stem Cell Transplant Mentor: Louis Constine, M.D.
Christine Zanghi Project Title: Anticoagulation in the era of Heparin Induced Thrombocytopenia: Combination Therapy with Bivalirudin and Fondaparinux Versus Conventional Heparin During Cardiopulmonary Bypass Mentor: Michael Eaton, M.D.
Past MD/PhD Trainees
Youngsun Cho Project Title: Sources of Dopamine Input to the Amygdala-an Anatomic and Neurochemical Focus of Mood Disorders Mentor: Julie Fudge, M.D.
Katherine Eisenberg Project Title: Lead Exposure in Refugee Children Mentor: Edwin Van Wijngaarden, Ph.D.
Laura Fornarola Project Title: Determining the role of the prolyl isomerase Pin1 in neuronal programmed cell death Mentor: Robert Freeman, Ph.D.
Natalia Golub Project Title: Longitudinal Health Outcomes in Former Refugees Mentor: Diana Fernandez, M.D., M.P.H., Ph.D.
Michael Jacob Project Title: Cognitive Influences on Perception: Studies in the neurophysiology of human aging and single cell dynamics Mentor: Charles Duffy, M.D.
Susan Lee Project Title: Neural basis of audiovisual integration during language comprehension in autism Mentor: Loisa Bennetto, Ph.D.
Kevin Makino Project Title: An Exploration of the Role of Public Health Insurance in Moderating the Effects of Low Family Income on Children's Educational Success Mentor: Bruce Friedman, Ph.D.
Kofi Mensah Project Title: Myelopoiesis in Erosive vs. Non-erosive Inflammatory Arthritis Mentor: Edward Schwarz, Ph.D.
Phillip Rappold Project Title: The Therapeutic Potential of Targeting Mitochondrial Fission/Fusion Pathways in Parkinson's Disease Mentor: Kim Tieu, Ph.D.
Christopher Richardson Project Title: B-Cell Tolerance and Immunology Mentor: Ignacio Sanz, M.D.
Adam Simning Project Title: Mental Illness among Older Adult Public Housing Residents Mentor: Edwin VanWijngaarden, Ph.D.
Mercedes Szpunar Project Title: The Effects of Stress on Breast Cancer: Beta-Adrenergic Receptor Stimulation of Cell Lines In Vitro and In Vivo Mentor: Edward Brown
Helen Wei Project Title: Astrocyte regulation of the cerebral microcirculation Mentors: Maiken Nedergaard, M.D., DM.Sc. and Edward Vates, M.D., Ph.D.
A Perception-Based Curricular Review on the K to 12 HUMSS Strand Curriculum
- November 2020
- IAFOR Journal of Education 8(4):25-44

- University of the Philippines Cebu
Abstract and Figures

Discover the world's research
- 25+ million members
- 160+ million publication pages
- 2.3+ billion citations
- Reneth Mae A. Diaz

- Cynthia Lubiton

- Ahmad Mazli bin Muhammad
- Aini Akmar binti Mohd Kasim

- Dolgun Aslan

- ShinSangKeun
- رضا فزونی شره جینی
- سیروس اسدیان
- National Research Council
- Division of Behavioral and Social Sciences and Education
- Board on Science Education
- Committee on a Conceptual Framework for New Science Education Standards

- Recruit researchers
- Join for free
- Login Email Tip: Most researchers use their institutional email address as their ResearchGate login Password Forgot password? Keep me logged in Log in or Continue with Google Welcome back! Please log in. Email · Hint Tip: Most researchers use their institutional email address as their ResearchGate login Password Forgot password? Keep me logged in Log in or Continue with Google No account? Sign up
Click through the PLOS taxonomy to find articles in your field.
For more information about PLOS Subject Areas, click here .
Loading metrics
Open Access
Peer-reviewed
Research Article
Academic Tracker: Software for tracking and reporting publications associated with authors and grants
Roles Methodology, Software, Validation, Writing – original draft, Writing – review & editing
Affiliation Superfund Research Center, University of Kentucky, Lexington, KY, United States of America
Roles Conceptualization, Methodology, Writing – review & editing
Affiliations Superfund Research Center, University of Kentucky, Lexington, KY, United States of America, Department of Computer Science (Data Science Program), University of Kentucky, Lexington, KY, United States of America, Markey Cancer Center, University of Kentucky, Lexington, KY, United States of America
Roles Conceptualization, Funding acquisition, Methodology, Project administration, Supervision, Writing – original draft, Writing – review & editing
* E-mail: [email protected]
Affiliations Superfund Research Center, University of Kentucky, Lexington, KY, United States of America, Markey Cancer Center, University of Kentucky, Lexington, KY, United States of America, Department of Molecular and Cellular Biochemistry, University of Kentucky, Lexington, KY, United States of America, Institute for Biomedical Informatics, University of Kentucky, Lexington, KY, United States of America, Center for Clinical and Translational Science, University of Kentucky, Lexington, KY, United States of America
- P. Travis Thompson,
- Christian D. Powell,
- Hunter N. B. Moseley

- Published: November 18, 2022
- https://doi.org/10.1371/journal.pone.0277834
- Peer Review
- Reader Comments
In recent years, United States federal funding agencies, including the National Institutes of Health (NIH) and the National Science Foundation (NSF), have implemented public access policies to make research supported by funding from these federal agencies freely available to the public. Enforcement is primarily through annual and final reports submitted to these funding agencies, where all peer-reviewed publications must be registered through the appropriate mechanism as required by the specific federal funding agency. Unreported and/or incorrectly reported papers can result in delayed acceptance of annual and final reports and even funding delays for current and new research grants. So, it’s important to make sure every peer-reviewed publication is reported properly and in a timely manner. For large collaborative research efforts, the tracking and proper registration of peer-reviewed publications along with generation of accurate annual and final reports can create a large administrative burden. With large collaborative teams, it is easy for these administrative tasks to be overlooked, forgotten, or lost in the shuffle. In order to help with this reporting burden, we have developed the Academic Tracker software package, implemented in the Python 3 programming language and supporting Linux, Windows, and Mac operating systems. Academic Tracker helps with publication tracking and reporting by comprehensively searching major peer-reviewed publication tracking web portals, including PubMed, Crossref, ORCID, and Google Scholar, given a list of authors. Academic Tracker provides highly customizable reporting templates so information about the resulting publications is easily transformed into appropriate formats for tracking and reporting purposes. The source code and extensive documentation is hosted on GitHub ( https://moseleybioinformaticslab.github.io/academic_tracker/ ) and is also available on the Python Package Index ( https://pypi.org/project/academic_tracker ) for easy installation.
Citation: Thompson PT, Powell CD, Moseley HNB (2022) Academic Tracker: Software for tracking and reporting publications associated with authors and grants. PLoS ONE 17(11): e0277834. https://doi.org/10.1371/journal.pone.0277834
Editor: Yuji Zhang, University of Maryland Baltimore, UNITED STATES
Received: April 1, 2022; Accepted: November 3, 2022; Published: November 18, 2022
Copyright: © 2022 Thompson et al. This is an open access article distributed under the terms of the Creative Commons Attribution License , which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Data Availability: All relevant data are located at: https://doi.org/10.6084/m9.figshare.19412165 .
Funding: This work was supported in part by grants NSF 2020026 (PI Moseley - HNBM), NIH P42 ES007380 (PI Pennell; co-I HNBM) via the Data Management and Analysis Core (DMAC), and NIH U54 TR001998-05A1 (PI Kern; co-I HNBM). There was no additional external funding received for this study. The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Competing interests: The authors have declared that no competing interests exist.
Introduction
Since 2008, the United States government has passed laws and issued directives to promote public access to peer-reviewed publications resulting from federal funding. These requirements started with Division G, Title II Section 218 of the Public Law (PL) 110–161 also known as the Consolidated Appropriations Act of 2008 [ 1 ], which directed the National Institutes for Health (NIH) to require all peer-reviewed publications supported by NIH funds to be electronically submitted to PubMed [ 2 ] within 12 months of the official date of publication [ 3 ]. Second in 2013, the White House Office of Science & Technology Policy (OSTP) mandated that all federal agencies with research and development budgets over $100 million to develop public access plans for research publications and data resulting from grants provided by these federal agencies [ 4 ]. Shortly thereafter in 2014, the US Congress passed the FY 2014 Omnibus Appropriations Act [ 5 ], which required federal agencies under Labor, Health and Human Services, and Education with research budgets of $100 million or more to provide public online access to peer-reviewed publications within 12 months of the official data of publication [ 6 ]. To comply with federal law, both NIH and NSF have implemented public access policies to make research supported by funding from these federal agencies freely available to the public. The enforcement of these policies typically occurs during the submission of annual and final reporting process for funded grants from NIH and NSF. In these reports, all peer-reviewed publications must be registered through the required mechanism by the specific federal funding agency. For NIH, peer-reviewed publications must be registered with PubMed Central and have a PubMed Central ID (PMCID). For NSF, peer-reviewed publications must be submitted to the NSF Public Access Repository (NSF-PAR) via Research.gov in the form of an archival PDF (PDF/A) [ 7 ]. Unreported and/or incorrectly reported papers can result in delayed acceptance of annual and final reports and funding delays for current and new research grants. Therefore, timely reporting of every peer-reviewed publication is required. For large collaborative research efforts involving large research teams or even multiple research teams, the tracking and proper registration of peer-reviewed publications along with generation of accurate annual and final reports can create a large administrative burden. With large collaborative teams, it is easy for these administrative tasks to be overlooked, forgotten, or lost in the shuffle.
In an effort to help researchers and their minders stay up-to-date with the reporting of peer-reviewed publications, we created the Academic Tracker software package. Written in the Python 3 programming language, Academic Tracker comprehensively searches major peer-reviewed publication tracking web portals, gathering relevant publications and useful tracking characteristics, for example, an indication of whether the publication has been reported to the NIH (is on PubMed), needs to be reported (is associated with an NIH grant), or satisfies the NIH’s requirements to have a PMCID. It has the ability to search PubMed [ 2 ], ORCID [ 8 ], Google Scholar [ 9 ], and Crossref [ 10 ], given a list of authors and/or author IDs. Academic Tracker provides highly customizable reporting templates so information about the resulting publications is easily transformed into appropriate formats for tracking and reporting purposes.
ORCID (Open Researcher and Contributor ID) is a non-profit organization dedicated to uniquely identifying individuals who participate in research [ 8 ]. Once an author is registered, ORCID provides a unique ID that can be used to associate an author with their publications. These associations can be easily accessed from the ORCID website or through their application programming interface (API). Google Scholar is a search engine for scholarly literature with similar API search facilities to ORCID [ 9 ]. Authors can create profiles on Google Scholar, which Google Scholar uses to automatically associate publications with. Crossref is a non-profit association with both commercial and non-profit publisher members with a primary purpose of enabling cross-publishing citation linking [ 10 ]. Crossref’s stated goal is to make “research objects easy to find, cite, link, assess, and reuse.” For the purposes of Academic Tracker, Crossref serves as a database with an easily accessible API for finding relevant publications.
Academic Tracker has three main use-cases and one supportive use-case. The first main use-case searches the aforementioned web portals for publications, given a list of authors. The second main use-case searches PubMed and Crossref for publication information, given a list of publication citations. Neither ORDID nor Google Scholar can be searched for specific publication information directly. ORCID is organized around author profiles and not publications themselves and does not provide a search option by publication characteristics. Google Scholar cannot be searched by specific publication characteristics, because Google Scholar has limited the repetitive programmatic use of their web service in this way. However, Google Scholar does allow repetitive programmatic search by author profile ID. The third main use-case finds collaborators given a list of authors. This is similar to the first use-case, but focuses on compiling the co-authors from the publications rather than the publications themselves. The fourth supportive use-case searches ORCID or Google Scholar for authors’ unique IDs for these sources, given a list of authors.
The main output from the three main use-cases is a Javascript Object Notation (JSON) file containing information about each publication found. Other outputs vary on user settings. Customizable summary and project reports can be created with an option of emailing them as attachments. The collaborator report of the third use-case is also customizable. All emails are also copied into a JSON file. A configuration JSON file is needed as part of the input to Academic Tracker and the fourth supportive use-case will update this file with the information found during the search. A use-case diagram for Academic Tracker is shown in Fig 1 .
- PPT PowerPoint slide
- PNG larger image
- TIFF original image
The first and third use-cases, publication search and collaborator search, are illustrated via the “Publication Search by Author” option. The second use-case, publication information, is illustrated via the “Publication Search by Reference” option. The supporting use-case, ORCID ID and Google Scholar ID searches, are illustrated by the “Unique ID Search” option.
https://doi.org/10.1371/journal.pone.0277834.g001
3 rd party packages
Academic Tracker leverages many third-party Python libraries and packages to accomplish its major tasks. Academic Tracker uses the docopt library to implement a command line interface (CLI) from a Python docstring description. Next, Academic Tracker uses the jsonschema library to validate user JSON input against an expected schema, which is also in JSON format. JSON Schema is an independently developed vocabulary or framework created for the purpose of validating and annotating JSON. Other developers have implemented the vocabulary in several languages, and the jsonschema library is the Python language implementation. The specific schema used in Academic Tracker are in the Validation_Schemas directory of the supplemental materials. Academic Tracker also uses four different packages to query data sources for publications. Specifically, Academic Tracker uses the pymed, habanero, orcid, and scholarly libraries to query PubMed, Crossref, ORCID, and Google Scholar, respectively. For the second use-case, Academic Tracker uses the requests library to make HTTP requests and the beautifulsoup4 library to parse HTML in the pulled web pages given as the reference file. Next, Academic Tracker uses the fuzzywuzzy library to fuzzy match publication titles, which is necessary because publications do not have a universal unique identifier. For general file input/output, Academic Tracker uses several packages, including: i) the python-docx library to read Microsoft Word files, specifically for the reference file input; ii) the pandas library to read and write tabular data, specifically to read in author data and write out reports; and iii) indirectly the openpyxl library, which is used by pandas to write Excel files. In order to comprehensively compare publication information across different runs to see if any information has changed, Academic Tracker uses the deepdiff library. A list of packages and their versions are in Table 1 .
https://doi.org/10.1371/journal.pone.0277834.t001
Although there are 3 main use-cases and 1 supportive use-case, Academic Tracker has 2 main commands and 6 supporting commands ( Table 2 ). The first and third main use-cases are handled by the author_search command, while the second main use-case is handled by the reference_search command. The supportive use-case is handled by the find_ORCID and find_Google_Scholar commands. The remaining four commands help users experiment with the tokenization and reporting systems in Academic Tracker and make it a little easier to convert author information into JSON format. The commands are listed in Table 2 . The input and output files for each command are further described in Table 3 .
https://doi.org/10.1371/journal.pone.0277834.t002
https://doi.org/10.1371/journal.pone.0277834.t003
Module description
Although Academic Tracker is primarily designed to be a command line tool, it does provide an equivalent API, which can be utilized if so desired. The CLI and highest-level API for each command are implemented in the __main__.py file, but other submodules break down the steps into smaller pieces. Utilizing the API, reference_search and author_search are almost completely separated into their own submodules. The athr_srch_modularized.py submodule compartmentalizes the steps of author_search, while the athr_srch_webio.py and athr_srch_emails_and_reports.py submodules contain the functions to interface with the internet and generate reports and emails respectively. reference_search is organized the same way with the ref_srch_modularized.py, ref_srch_webio.py, and ref_srch_emails_and_reports.py submodules. The user_input_checking.py submodule contains the functions to validate user input for errors, and the tracker_schema.py submodule works in tandem with it to store the JSON schema being used for validation. The fileio.py submodule contains all the functions for reading and writing files. The webio.py submodule contains functions to interface with the internet that are more general purpose or common to multiple commands. It is where the functions to interface with the internet for find_ORCID and find_Google_Scholar are. The helper_functions.py submodule contains functions with common operations across all commands that don’t classify well into any other submodule, such as regex operations and data transformation. The citation_parsing.py submodule contains all the functions used to tokenize the reference sources for reference_search. Table 4 lists the submodules of Academic Tracker, and Fig 2 shows a module diagram.
Submodule and module dependencies are illustrated by connecting lines, except for helper_functions which is utilized by most other submodules.
https://doi.org/10.1371/journal.pone.0277834.g002
https://doi.org/10.1371/journal.pone.0277834.t004
The Academic Tracker package was originally developed in a Linux operating system (OS) environment, but has been directly tested on Linux, Windows, and MacOS operating systems. All use-cases have been tested on these operating systems; however, Academic Tracker relies on sendmail or an emulator being installed and configured on the machine for its email functionality. In addition, each submodule includes unit-tests that test all critical functions of the submodule. Every function in every module is tested to make sure it gives the expected output when it should and errors when it should. All requests to web portals are replaced with mock data. The user_input_checking.py submodule has the largest number of tests, since it tests several error states for each element of the input JSON files. Every command line option is tested, for example, silent and not searching ORCID options. Various ways of creating reports are also tested, such as creating a tabular report versus a text report, Excel versus CSV format, and renaming the report from the default name. Several different citation styles and sources are also tested to make sure they are tokenized correctly, such as MEDLINE, a MyNCBI bibliography URL, and an NSF Award page.
Academic Tracker can be utilized in many different ways and was designed with a great deal of flexibility, anticipating users’ desire to use it in unpredictable ways. However, the three main and one supportive use-case are presented here. Note that the figures here are general examples with mostly dummy data. There are full examples with real data and run commands in the supplemental materials (Example_Runs subdirectory). The first main use-case involves searching for publications given author information. Fig 3 shows an example input configuration JSON file, the command line for its execution, the API execution equivalent, and the resulting output files. Fig 4 shows the contents of these resulting output files. Authors without unique ORCID or Google Scholar IDs are identified by matching first name, last name, and at least one affiliation.
Example configuration file, command-line execution, API execution, and file output of the author_search for publications use-case shown.
https://doi.org/10.1371/journal.pone.0277834.g003
Example JSON publications output and plain-text summary report from the author_search for publications use-case shown.
https://doi.org/10.1371/journal.pone.0277834.g004
The second main use-case involves looking for publications based on a given reference. Fig 5 shows an example input configuration JSON file, the command line for its execution, the API execution equivalent, and the resulting output files. Figs 6 and 7 show the contents of the resulting output files.
Example configuration file, reference file, command-line execution, API execution, and file output of the reference_search use-case shown.
https://doi.org/10.1371/journal.pone.0277834.g005
https://doi.org/10.1371/journal.pone.0277834.g006
https://doi.org/10.1371/journal.pone.0277834.g007
The third use-case is basically identical to the first, but a collaborator report attribute needs to be added to an author. Fig 8 is essentially the same as Fig 3 , but with a collaborator report attribute added to Author1 and the report in the output directory. Fig 4 already shows the contents of the publications JSON and summary report. Table 5 shows the contents of the resulting collaborator report table.
Example configuration file, command-line execution, API execution, and file output of the author_search for collaborators use-case shown.
https://doi.org/10.1371/journal.pone.0277834.g008
https://doi.org/10.1371/journal.pone.0277834.t005
The supportive use-case is broken into 2 commands: find_ORCID for finding ORCID IDs and find_Google_Scholar for finding Google Scholar IDs. Fig 9 shows an example input configuration JSON, how to accomplish this using the command line and API, and the resulting output files for finding ORCID IDs. Fig 10 shows the contents of the resulting configuration JSON file. Figs 11 and 12 are the same as Figs 9 and 10 but for finding Google Scholar IDs.
Example configuration file, command-line execution, API execution, and file output of the author ORCID ID search use-case shown.
https://doi.org/10.1371/journal.pone.0277834.g009
https://doi.org/10.1371/journal.pone.0277834.g010
Example configuration file, command-line execution, API execution, and file output of the author Google Scholar ID search use-case shown.
https://doi.org/10.1371/journal.pone.0277834.g011
https://doi.org/10.1371/journal.pone.0277834.g012
Discussion and conclusions
Academic Tracker is a useful tool for querying major scientific publication web portals for publications, given a list of authors or references and for creating highly customizable reports from the list of publications found. The software package provides assistance in repetitive tracking and reporting of peer-reviewed publications associated with specific authors, projects, and grants. Specifically, the JSON configuration file supports batch execution, directing Academic Tracker to perform multiple related author searches and report generations. The JSON configuration file has many optional parameters to customize searching and report generation, including a cutoff_year for searching. Academic Tracker is also designed for repetitive tracking by comparing current search results to prior search results to limit reporting to changes in publications detected and in publication attributes. Academic Tracker also provides facilities for generating lists of co-author collaborators, which has several uses in grant proposal submission. But given the number of major use-cases and versality of the software, there is some intellectual overhead required to initially setup the JSON configuration file and customize reports. Additional supportive commands are included to make learning and troubleshooting the tool easier for new users. Also, there is extensive documentation available to help with the learning curve: https://moseleybioinformaticslab.github.io/academic_tracker/
In addition, when installed via the Python package management system pip, a console script “academic_tracker” is created automatically for the user, providing easy access to the CLI.
While the package accesses multiple major peer-reviewed publication tracking web portals, it is fundamentally limited to the information provided by these web portals and must assume the information provided is accurate. One possibility is to download a PDF of the publication itself for analysis. However, this is pragmatically infeasible, since there is wide variation in how journals organize the splash page of their publications. One way to alleviate this issue is for journals to adopt a DOI extension like “.pdf” which would link directly to the PDF version of the publication, if the PDF version is accessible. This is similar to the versioning “.v#” DOI extension that FigShare uses to provide access each version of a public FigShare repository. If a practical way to directly access the PDF is implemented either by journals or the publication tracking web portals, we would extend Academic Tracker to utilize it. Still in its current implementation, we believe Academic Tracker can significantly reduce the stress and hassle of reporting publications to federal funding agencies, reducing the chance for accidental non-compliance and resulting delay in funding.
Acknowledgments
We also thank Jennifer Moore for feedback during the development of the report generation capabilities.
- View Article
- Google Scholar
- 5. Consolidated Appropriations Act, 2014, H.R. 3547, Editor. 2013: Congressional Record.

IMAGES
VIDEO
COMMENTS
Previous research shows academic track schools are more successful than non-academic track schools in teaching mathematics, reading and foreign languages. Reasons include a more favorable student composition and higher instructional quality. However, there is less evidence that between track differences are even large enough to differentially ...
The Tukey's post-hoc test revealed that students who selected an academic track at an all-academic school had higher academic self-concept in their last year of compulsory school than academic students in multilateral schools (MD = 0.22, SE = 0.05, p < .001), with a small to medium difference in terms of Cohen's d (d s = 0.
In light of the scarce amount of research on relationships between school-track-related stereotype awareness and academic outcomes, it seemed most appropriate to rely on theoretical considerations ...
Future research could substantiate that these track differences in teachers' behaviors lead to differences across time in the academic achievement and goals of early adolescents. Research suggests that students in standard classes are aware that teachers demonstrate more positive interactions with students in advanced classes ( Gilbert ...
academically tended to choose the academic track, while those who Keywords: Senior high school, Track preference, Determinants, G10 students. ... this research was grade 10 students; both male and female adolescents aged 15-17 years old, enrolled in the school year 2021-2022. Two schools from private and four schools from the public were
Academic tracking has long been a subject of debate due to its potential impact on educational equity, with students who are tracked highly receiving a higher quality education in comparison to students tracked lowly. These disparities in education quality may be affecting students' outcomes, as it has been demonstrated that the short-term academic outcomes of students, such as their grades ...
This study aims to assess some selected factors associated with of preferences of senior high school track. that are commonly adopted by the Grade 10 students of a secondary natio nal high school ...
Previous research shows academic track schools are more successful than non-academic track schools in teaching mathematics, reading and foreign languages. Reasons include a more favorable student ...
Academic tracking is a common feature of school organization, but it produces inequalities in student outcomes. ... Same school, separate worlds: A sociocultural study of identity, resistance, and negotiation in a rural, lower track science classroom. Journal of Research in Science Teaching, 38, 574-598. Crossref. Web of Science. Google Scholar.
CORRELATES OF STUDENTS PREFERENCE ON TVL TRACK AND ACADEMIC ENGAGEMENT. December 2018. Thesis for: BACHELOR OF SCIENCE IN TECHNOLOGY TEACHER EDUCATION MAJOR IN INDUSTRIAL TECHNOLOGY. Advisor: SIR ...
There are many influences that affect the preferences of grade 10 students in choosing a track to proceed to senior high school. Likewise, this study aims to identify influence of preference of a Senior High School track that is commonly encountered by the Grade 10 students in terms of Gender, Socio-Economic Status, Average academic grades, nature of parent's occupation; and, strand and the ...
There are many influences that affect the preferences of grade 10 students in choosing a track to proceed to senior high school. Likewise, this study aims to identify influence of preference of a Senior High School track that is commonly encountered by the Grade 10 students in terms of Gender, Socio-Economic Status, Average academic grades, nature of parent's occupation; and, strand and the ...
The Research Whisperer is dedicated to the topic of doing research in academia. We talk about finding funding, research culture, and building academic track-records. This blog is managed by Tseen Khoo and Jonathan O'Donnell.
The data were collected from Grade 12 Senior High School Academic Track students with the use of the English Speaking Attitude Questionnaire (ESAQ). ... especially attitude, should be considered in language research. Senior High School students are expected to have better English language proficiency, especially their oral communication ability
Today, they need proven track records that include conference presentations and peer-reviewed publications. They cannot wait for graduation to start building their Curriculum Vitae (CV), as competition for grants, post-doctorate positions, and jobs require that students are already active in their fields.
The goals of the Longitudinal Academic Tracks are to promote intellectual curiosity, appreciation of scholarly inquiry, inter-professional collaboration, and cura personalis. Longitudinal Academic Tracks currently offered in the School of Medicine: Diversity, Equity, & Inclusion in Medicine Track. Environmental Health and Medicine Track.
K to 12 ›. Academic Track. Sample Scheduling of Subjects Accountancy, Business and Management (ABM) Strand. Sample Scheduling of Subjects. Applied Economics. Business Ethics and Social Responsibility. Fundamentals of Accountancy, Business and Management 1. Fundamentals of Accountancy, Business and Management 2. Business Math.
From social science to computer science, X data can advance research objectives on topics as diverse as the global conversations happening on X. Help us design for your needs by taking part in our academic research panel. Your feedback will help shape our investments that serve the academic research community.
Clinical & Translational Science Institute / Education and Career / Academic Research Track (ART) Academic Research Track (ART) The Academic Research Track (ART) was a program for first, second and third year medical students who considered a research experience as part of their education or career. Jump to:
The data were collected from G rade 12 Senior High School Academic. Track students with the use of the English Speaking Attitude Questionnaire. (ESAQ). R esults show that both HumSS and ABM strand ...
What is a Research Track Record? M. Polonsky Australasian Marketing Journal 16 (2), 2008 67 Commentaries from the Research Network Collaboration Special Session ANZMAC 2006 Introduction1. The evaluation of academic research performance is a difficult question and can be evaluated in a number of different ways.
Academic, Technical-Vocational Livelihood, Sports, and Arts and Design Tracks (Shahani, 2015). Among the strands under the Academic Track is the Humanities and Social Sciences
For large collaborative research efforts, the tracking and proper registration of peer-reviewed publications along with generation of accurate annual and final reports can create a large administrative burden. ... Windows, and Mac operating systems. Academic Tracker helps with publication tracking and reporting by comprehensively searching ...