- Open access
- Published: 22 April 2020

Research and trends in STEM education: a systematic analysis of publicly funded projects
- Yeping Li 1 ,
- Ke Wang 2 ,
- Yu Xiao 1 ,
- Jeffrey E. Froyd 3 &
- Sandra B. Nite 1
International Journal of STEM Education volume 7 , Article number: 17 ( 2020 ) Cite this article
17k Accesses
32 Citations
7 Altmetric
Metrics details
Taking publicly funded projects in STEM education as a special lens, we aimed to learn about research and trends in STEM education. We identified a total of 127 projects funded by the Institute of Education Sciences (IES) of the US Department of Education from 2003 to 2019. Both the number of funded projects in STEM education and their funding amounts were high, although there were considerable fluctuations over the years. The number of projects with multiple principal investigators increased over time. The project duration was typically in the range of 3–4 years, and the goals of these projects were mostly categorized as “development and innovation” or “efficacy and replication.” The majority of the 127 projects focused on individual STEM disciplines, especially mathematics. The findings, based on IES-funded projects, provided a glimpse of the research input and trends in STEM education in the USA, with possible implications for developing STEM education research in other education systems around the world.
Introduction
The rapid development of science, technology, engineering, and mathematics (STEM) education and research since the beginning of this century has benefited from strong, ongoing support from many different entities, including government agencies, professional organizations, industries, and education institutions (Li, 2014 ). Typically, studies that summarized the status of research in STEM education have used publications as the unit of their analyses (e.g., Li et al., 2019 ; Li et al., 2020 ; Margot & Kettler, 2019 ; Minichiello et al., 2018 ; Otten, Van den Heuvel-Panhuizen, & Veldhuis, 2019 ; Schreffler et al., 2019 ). Another approach, which has been used less frequently, is to study research funding. Although not all research publications were generated from funded projects and not all funded projects have been equally productive, as measured by publications, research funding and publications present two different, but related perspectives on the state of research in STEM education. Our review focuses on research funding.
Types of funding support to education research
There are different types of sources and mechanisms in place to allocate, administer, distribute, and manage funding support to education. In general, there are two sources of funding: public and private.
Public funding sources are commonly government agencies that support education program development and training, project evaluation, and research. For example, multiple state and federal agencies in the USA provide and manage funding support to education research, programs and training, including the US Department of Education (ED), the National Science Foundation (NSF), and the National Endowment for the Humanities—Division of Education Programs. Researchers seeking support from public funding sources often submit proposals that are vetted through a well-structured peer-review process. The process is competitive, and the decision to fund a project validates both its importance and alignment with the funding agency’s development agenda. Changes in the agencies’ agendas and funding priorities can reflect governmental intentions and priorities for education and research.
Private funding sources have played a very important role in supporting education programs and research with a long history. Some private funding sources in the USA can be sizeable, such as the Bill & Melinda Gates Foundation ( https://www.gatesfoundation.org ), while many also have specific foci, such as the Howard Hughes Medical Institute ( https://www.hhmi.org ) that is dedicated to advancing science through research and science education. At the same time, private funding sources often have their own development agendas, flexibility in deciding funding priorities, and specific mechanisms in making funding decisions, including how funds can be used, distributed, and managed. Indeed, private funding sources differ from public funding sources in many ways. Given many special features associated with private funding sources, including the lack of transparency, we chose to examine projects that were supported by public funding sources in this review.
Approaches to examining public research funding support
One approach to studying public research funding support to STEM education would be to examine requests-for-proposals (RFPs) issued by different government agencies. However, those RFPs tend to provide guidelines, which are not sufficiently concrete to learn about specific research that is funded. In contrast, reviewing those projects selected for funding can provide more detailed information on research activity. Figure 1 shows a flowchart of research activity and distinguishes how funded projects and publications might provide different perspectives on research. In this review, we focus on the bolded portion of the flowchart, i.e., projects funded to promote STEM education.

A general flowchart of RFPs to publications
Current review
Why focus on research funding in the usa.
Recent reviews of journal publications in STEM education have consistently revealed that scholars in the USA played a leading role in producing and promoting scholarship in STEM education, with about 75% of authorship credits for all publications in STEM education either in the International Journal of STEM Education alone from 2014 to 2018 (Li et al., 2019 ) or in 36 selected journals published from 2000 to 2018 (Li et al., 2020 ). The strong scholarship development in the USA is likely due to a research environment that is well supported and conducive to high research output. Studying public funding support for STEM education research in the USA will provide information on trends and patterns, which will be valuable both in the USA and in other countries.
The context of policy and public funding support to STEM education in the USA
The tremendous development of STEM education in the USA over the past decades has benefited greatly from both national policies and strong funding support from the US governmental agencies as well as private funding sources. Federal funding for research and development in science, mathematics, technology, and engineering-related education in the USA was restarted in the late 1980s, in the latter years of the Reagan administration, which had earlier halted funding. In recent years, the federal government has strongly supported STEM education research and development. For example, the Obama administration in the USA (The White House, 2009 ) launched the “Educate to Innovate” campaign in November 2009 for excellence in STEM education as a national priority, with over 260 million USD in financial and in-kind support commitment. The Trump administration has continued to emphasize STEM education. For example, President Trump signed a memorandum in 2017 to direct ED to spend 200 million USD per year on competitive grants promoting STEM (The White House, 2017 ). In response, ED awarded 279 million USD in STEM discretionary grants in Fiscal Year 2018 (US Department of Education, 2018 ). The Trump administration took a step further to release a report in December 2018 detailing its five-year strategic plan of boosting STEM education in the USA (The White House, 2018 ). The strategic plan envisions that “All Americans will have lifelong access to high-quality STEM education and the USA will be the global leader in STEM literacy, innovation, and employment.” (Committee on STEM Education, 2018 , p. 1). Consistently, current Secretory of Education DeVos in the Trump administration has taken STEM as a centerpiece of her comprehensive education agenda (see https://www.ed.gov/stem ). The consistency in national policies and public funding support shows that STEM education continues to be a strategic priority in the USA.
Among many federal agencies that funded STEM education programs, the ED and NSF have functioned as two primary agencies. For ED, the Institute of Education Sciences (Institute of Education Sciences (IES), n.d. , see https://ies.ed.gov/aboutus/ ) was created by the Education Sciences Reform Act of 2002 as its statistics, research, and evaluation arm. ED’s support to STEM education research has been mainly administered and managed by IES since 2003. In contrast to the focus of ED on education, NSF (see https://www.nsf.gov/about/ ) was created by Congress in 1950 to support basic research in many fields such as mathematics, computer sciences, and social sciences. Education and Human Resources is one of its seven directorates that provides important funding support to STEM education programs and research. In addition to these two federal agencies, some other federal agencies also provide funding support to STEM education programs and research from time to time.
Any study of public funding support to STEM education research in the USA would need to limit its scope, given the complexity of various public funding sources available in the system, the ambiguity associated with the meaning of STEM education across different federal agencies (Li et al., 2020 ), and the number of programs that have funded STEM education research over the years. For the purpose of this review, we have chosen to focus on the projects in STEM education funded by IES.
Research questions
Given the preceding research approach decision to focus on research projects funded by IES, we generated the following questions:
What were the number of projects, total project funding, and the average funding per project from 2003 to 2019 in STEM education research?
What were the trends of having single versus multiple principal investigator(s) in STEM education?
What were the types of awardees of the projects?
What were the participant populations in the projects?
What were the types of projects in terms of goals for program development and research in STEM education?
What were the disciplinary foci of the projects?
What research methods did projects tend to use in conducting STEM education research?
Based on the above discussion to focus on funding support from IES, we first specified the time period, and then searched the IES website to identify STEM education research projects funded by IES within the specified time period.
Time period
As discussed above, IES was established in 2002 and it did not start to administer and manage research funding support for ED until 2003. Therefore, we considered IES funded projects from 2003 to the end of 2019.
Searching and identifying IES funded projects in STEM education
Given the diverse perspectives about STEM education across different agencies and researchers (Li et al., 2020 ), we did not discuss and define the meaning of STEM education. Instead, we used the process described in the following paragraph to identify STEM education research projects funded by IES.
On the publicly accessible IES website ( https://ies.ed.gov ), one menu item is “FUNDING OPPORTUNITIES”, and there is a list of choices within this menu item. One choice is “SEARCH FUNDED RESEARCH GRANTS AND CONTRACTS.” On this web search page, we can choose “Program” under “ADDITIONAL SEARCH OPTIONS.” There are two program categories related to STEM under the option of “Program.” One is “Science, Technology, Engineering, and Mathematics (STEM) Education” under one large category of “Education Research” and the other is “Science, Technology, Engineering, and Mathematics” under another large category of “Special Education Research.” We searched for funded projects under these two program categories, and the process returned 98 funded projects in “Science, Technology, Engineering, and Mathematics (STEM) Education” under “Education Research” and 29 funded projects in “Science, Technology, Engineering, and Mathematics” under “Special Education Research,” for a total of 127 funded projects in these two programs designated for STEM education by IES Footnote 1 .
Data analysis
To address questions 1, 2, 3, and 4, we collected the following information about these projects identified using above procedure: amount of funding, years of duration, information about the PI, types of awardees that received and administered the funding (i.e., university versus those non-university including non-profit organization such as WestEd, Educational Testing Service), and projects’ foci on school level and participants. When a project’s coverage went beyond one category, the project was then coded in terms of its actual number of categories being covered. For example, we used the five categories to classify project’s participants: Pre–K, grades 1–4, grades 5–8, grades 9–12, and adult. If a funded project involved participants from Pre-school to grade 8, then we coded the project as having participants in three categories: Pre-K, grades 1–4, and grades 5–8.
To address question 5, we analyzed projects based on goal classifications from IES. IES followed the classification of research types that was produced through a joint effort between IES and NSF in 2013 (Institute of Education Sciences (IES) and National Science Foundation (NSF), 2013 ). The effort specified six types of research that provide guidance on the goals and level of funding support: foundational research, early-stage or exploratory research, design and development research, efficacy research, effectiveness research, and scale-up research. Related to these types, IES classified goals for funded projects: development and innovation, efficacy and replication, exploration, measurement, and scale-up evaluation, as described on the IES website.
To address question 6, we coded the disciplinary focus using the following five categories: mathematics, science, technology, engineering, and integrated (meaning an integration of any two or more of STEM disciplines). In some cases, we coded a project with multiple disciplinary foci into more than one category. The following are two project examples and how we coded them in terms of disciplinary foci:
The project of “A Randomized Controlled Study of the Effects of Intelligent Online Chemistry Tutors in Urban California School Districts” (2008, https://ies.ed.gov/funding/grantsearch/details.asp?ID=601 ) was to test the efficacy of the Quantum Chemistry Tutors, a suite of computer-based cognitive tutors that are designed to give individual tutoring to high school students on 12 chemistry topics. Therefore, we coded this project as having three categories of disciplinary foci: science because it was chemistry, technology because it applied instructional technology, and integrated because it integrated two or more of STEM disciplines.
The project of “Applications of Intelligent Tutoring Systems (ITS) to Improve the Skill Levels of Students with Deficiencies in Mathematics” (2009, https://ies.ed.gov/funding/grantsearch/details.asp?ID=827 ) was coded as having three categories of disciplinary foci: mathematics, technology because it used intelligent tutoring systems, and integrated because it integrated two or more of STEM disciplines.
To address question 7, all 127 projects were coded using a classification category system developed and used in a previous study (Wang et al., 2019 ). Specifically, each funded project was coded in terms of research type (experimental, interventional, longitudinal, single case, correlational) Footnote 2 , data collection method (interview, survey, observation, researcher designed tests, standardized tests, computer data Footnote 3 ), and data analysis method (descriptive statistics, ANOVA*, general regression, HLM, IRT, SEM, others) Footnote 4 . Based on a project description, specific method(s) were identified and coded following a procedure similar to what we used in a previous study (Wang et al., 2019 ). Two researchers coded each project’s description, and the agreement between them for all 127 projects was 88.2%. When method and disciplinary focus-coding discrepancies occurred, a final decision was reached after discussion.
Results and discussion
In the following sections, we report findings as corresponding to each of the seven research questions.
Question 1: the number of projects, total funding, and the average funding per project from 2003 to 2019
Figure 2 shows the distribution of funded projects over the years in each of the two program categories, “Education Research” and “Special Education Research,” as well as combined (i.e., “STEM” for projects funded under “Education Research,” “Special STEM” for projects funded under “Special Education Research,” and “Combined” for projects funded under both “Education Research” and “Special Education Research”). As Fig. 2 shows, the number of projects increased each year up to 2007, with STEM education projects started in 2003 under “Education Research” and in 2006 under “Special Education Research.” The number of projects in STEM under “Special Education Research” was generally less than those funded under the program category of “Education Research,” especially before 2011. There are noticeable decreases in combined project counts from 2009 to 2011 and from 2012 to 2014, before the number count increased again in 2015. We did not find a consistent pattern across the years from 2003 to 2019.
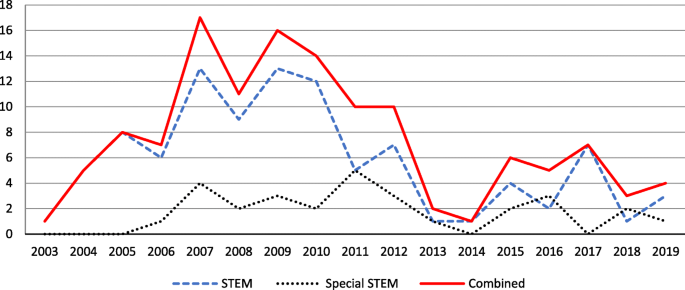
The distribution of STEM education projects over the years. (Note: STEM refers to projects funded under “Education Research,” Special STEM refers to projects funded under “Special Education Research,” and “Combined” refers to projects funded under both “Education Research” and “Special Education Research.” The same annotations are used in the rest of the figures.)
A similar trend can be observed in the total funding amount for STEM education research (see Fig. 3 ). The figure shows noticeably big year-to-year swings from 2003 to 2019, with the highest funding amount of more than 33 million USD in 2007 and the lowest amount of 2,698,900 USD in 2013 from these two program categories. Although it is possible that insufficient high-quality grant proposals were available in one particular year to receive funding, the funded amount and the number of projects (Fig. 2 ) provide insights about funding trends over the time period of the review.
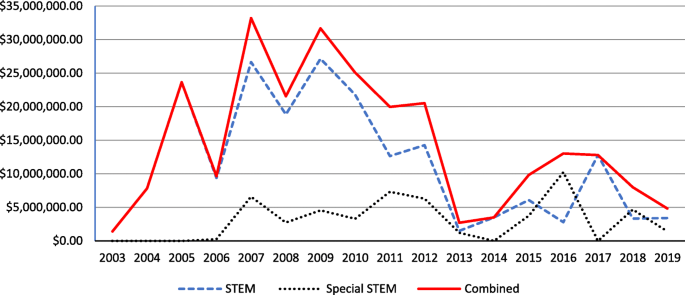
Annual funding totals
As there are diverse perspectives and foci about STEM education, we also wondered if STEM education research projects might be funded by IES but in program options other than those designated options of “Science, Technology, Engineering, and Mathematics (STEM) Education.” We found a total of 54 funded projects from 2007 to 2019, using the acronym “STEM” as a search term under the option of “SEARCH FUNDED RESEARCH GRANTS AND CONTRACTS” without any program category restriction. Only 2 (3.7%) out of these 54 projects were in the IES designated program options of STEM education in the category of “Education Research.” Further information about these 54 projects and related discussion can be found as additional notes at the end of this review.
Results from two different approaches to searching for IES-funded projects will likely raise questions about what kinds of projects were funded in the designated program option of “Science, Technology, Engineering, and Mathematics (STEM) Education,” if only two funded projects under this option contained the acronym “STEM” in a project’s title and/or description. We shall provide further information in the following sub-sections, especially when answering question 6 related to projects’ disciplinary focus.
Figure 4 illustrates the trend of average funding amount per project each year in STEM education research from 2003 to 2019. The average funding per project varied considerably in the program category “Special Education Research,” and no STEM projects were funded in 2014 and 2017 in this category. In contrast, average funding per project was generally within the range of 1,132,738 USD in 2019 to 3,475,975 USD in 2014 for the projects in the category of “Education Research” and also for project funding in the combined category.
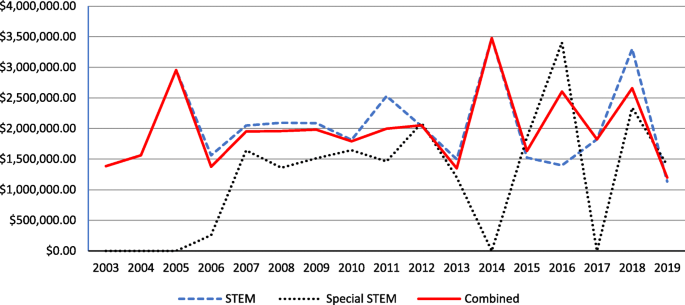
The trend of average funding amount per project funded each year in STEM education research
Figure 5 shows the number of projects in different funding amount categories (i.e., less than 1 million USD, 1–2 million USD, 2–3 million USD, 3 million USD or more). The majority of the 127 projects obtained funding of 1–2 million USD (77 projects, 60.6%), with 60 out of 98 projects (61.2%) under “Education Research” program and 17 out of 29 projects (58.6%) in the program category “Special Education Research.” The category with second most projects is funding of 3 million USD or more (21 projects, 16.5%), with 15 projects (15.3% of 98 projects) under “Education Research” and 6 projects (20.7% of 29 projects) under “Special Education Research.”
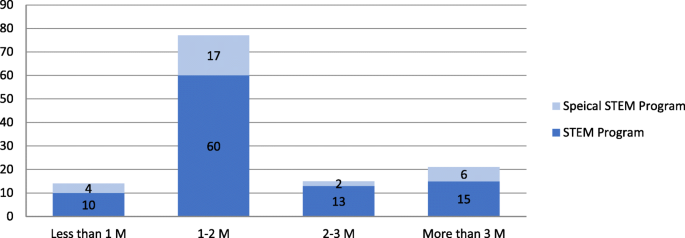
The number of projects in terms of total funding amount categories
Figure 6 shows the average amount of funding per project funded across these different funding amount and program categories. In general, the projects funded under “Education Research” tended to have a higher average amount than those funded under “Special Education Research,” except for those projects in the total funding amount category of “less than 1 million USD.” Considering all 127 funded projects, the average amount of funding was 1,960,826.3 USD per project.
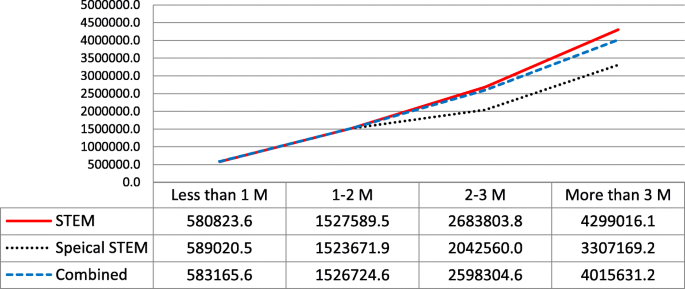
The average amount of funding per project across different total funding amount and program categories
Figure 7 shows that the vast majority of these 127 projects were 3- or 4-year projects. In particular, 59 (46.5%) projects were funded as 4-year projects, with 46 projects (46.9%) under “Education Research” and 13 projects (44.8%) under “Special Education Research.” This category is followed closely by 3-year projects (54 projects, 42.5%), with 41 projects (41.8%) under “Education Research” and 13 projects (44.8%) under “Special Education Research.”
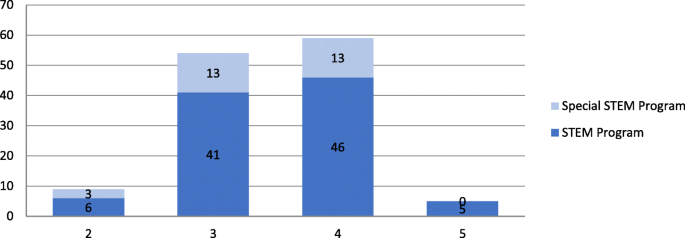
The number of projects in terms of years of project duration. (Note, 2: 2-year projects; 3: 3-year projects; 4: 4-year projects; 5: 5-year projects)
Question 2: trends of single versus multiple principal investigator(s) in STEM education
Figure 8 shows the distribution of projects over the years grouped by a single PI or multiple PIs where the program categories of “Education Research” and “Special Education Research” have been combined. The majority of projects before 2009 had a single PI, and the trend has been to have multiple PIs for STEM education research projects since 2009. The trend illustrates the increased emphases on collaboration in STEM education research, which is consistent with what we learned from a recent study of journal publications in STEM education (Li et al., 2020 ).
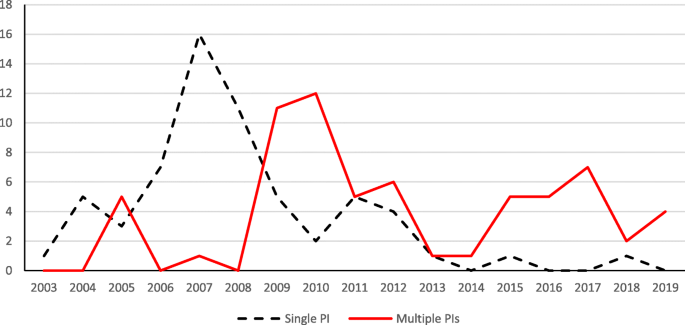
The distribution of projects with single versus multiple PIs over the years (combined)
Separating projects by program categories, Fig. 9 shows projects funded in the program category “Education Research.” The trends of single versus multiple PIs in Fig. 9 are similar to the trends shown in Fig. 8 for the combined programs. In addition, almost all projects in STEM education funded under this regular research program had multiple PIs since 2010.
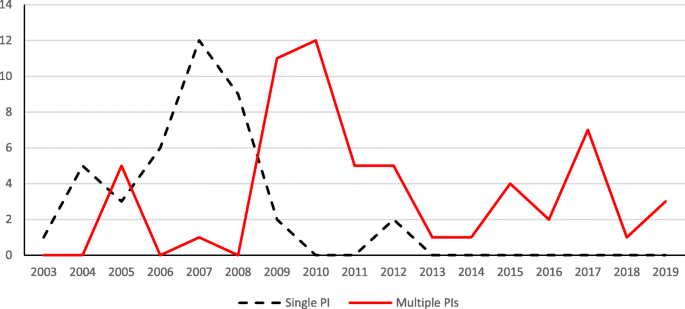
The distribution of projects with single versus multiple PIs over the years (in “Education Research” program)
Figure 10 shows projects funded in the category “Special Education Research.” The pattern in Fig. 10 , where very few projects funded under this category had multiple PIs before 2014, is quite different from the patterns in Figs. 8 and 9 . We did not learn if single PIs were appropriate for the nature of these projects. The trend started to change in 2015 as the number of projects with multiple PIs increased and the number of projects with single PIs declined.

The distribution of projects with single versus multiple PIs over the years (in “Special Education Research” program)
Question 3: types of awardees of these projects
Besides the information about the project’s PI, the nature of the awardees can help illustrate what types of entity or organization were interested in developing and carrying out STEM education research. Figure 11 shows that the university was the main type of awardee before 2012, with 80 (63.0%) projects awarded to universities from 2003 to 2019. At the same time, non-university entities received funding support for 47 (37.0%) projects and they seem to have become even more active and successful in obtaining research funding in STEM education over the past several years. The result suggests that diverse organizations develop and conduct STEM education research, another indicator of the importance of STEM education research.
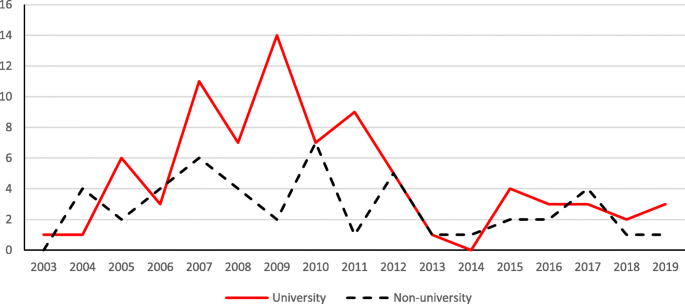
The distribution of projects funded to university versus non-university awardees over the years
Question 4: participant populations in the projects
Figure 12 indicates that the vast majority of projects were focused on student populations in preschool to grade 12. This is understandable as IES is the research funding arm of ED. Among those projects, middle school students were the participants in the most projects (70 projects), followed by student populations in elementary school (48 projects), and high school (38 projects). The adult population (including post-secondary students and teachers) was the participant group in 36 projects in a combined program count.
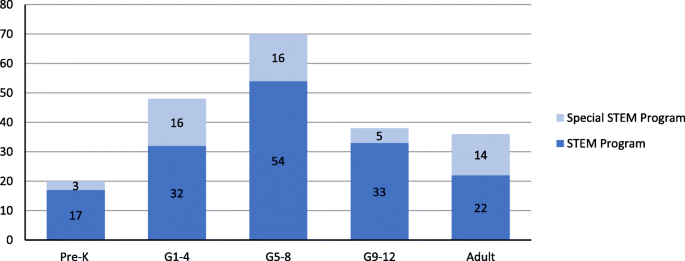
The number of projects in STEM education for different groups of participants (Note: Pre-K: preschool-kindergarten; G1–4: grades 1–4; G5–8: grades 5–8; G9–12: grades 9–12; adult: post-secondary students and teachers)
If we separate “Education Research” and “Special Education Research” programs, projects in the category “Special Education Research” focused on student populations in elementary and middle school most frequently, and then adult population. In contrast, projects in the category “Education Research” focused most frequently on middle school student population, followed by student populations in high school and elementary school.
Given the importance of funded research in special education Footnote 5 at IES, we considered projects focused on participants with disabilities. Figure 13 shows there were 28 projects in the category “Special Education Research” for participants with disabilities. There were also three such projects funded in the category “Education Research,” which together accounted for a total of 31 (24.4%) projects. In addition, some projects in the category “Education Research” focused on other participants, including 11 projects focused on ELL students (8.7%) projects and 37 projects focused on low SES students (29.1%).
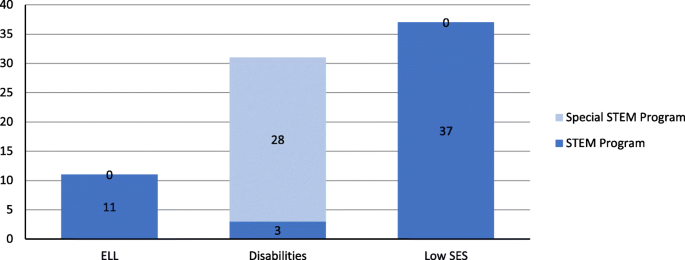
The number of funded projects in STEM education for three special participant populations (Note: ELL: English language learners, Low SES: low social-economic status)
Figure 14 shows the trend of projects in STEM education for special participant populations. Participant populations with ELL and/or Low SES gained much attention before 2011 among these projects. Participant populations with disabilities received relatively consistent attention in projects on STEM education over the years. Research on STEM education with special participant populations is important and much needed. However, related scholarship is still in an early development stage. Interested readers can find related publications in this journal (e.g., Schreffler et al., 2019 ) and other journals (e.g., Lee, 2014 ).
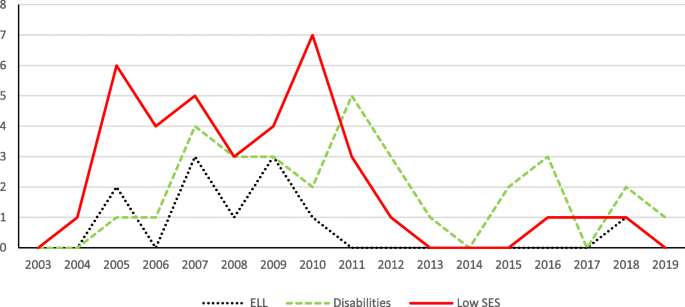
The distribution of projects in STEM education for special participant populations over the years
Question 5: types of projects in terms of goals for program development and research
Figure 15 shows that “development and innovation” was the most frequently funded type of project (58 projects, 45.7%), followed by “efficacy and replication” (34 projects, 26.8%), and “measurement” (21 projects, 16.5%). The pattern is consistent across “Education Research,” “Special Education Research,” and combined. However, it should be noted that all five projects with the goal of “scale-up evaluation” were in the category “Education Research” Footnote 6 and funding for these projects were large.
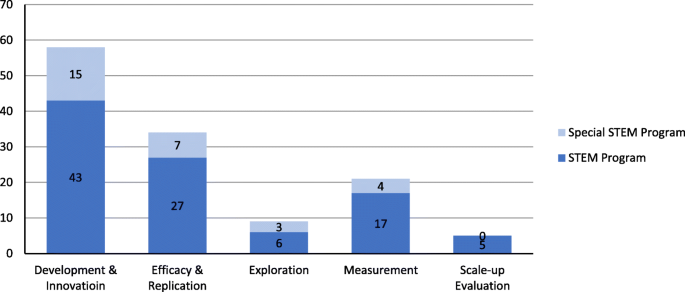
The number of projects in terms of the types of goals
Examining the types of projects longitudinally, Fig. 16 shows that while “development and innovation” and “efficacy and replication” types of projects were most frequently funded in the “Education Research” program, the types of projects being funded changed longitudinally. The number of “development and innovation” projects was noticeably fewer over the past several years. In contrast, the number of “measurement” projects and “efficacy and replication” projects became more dominant. The change might reflect a shift in research development and needs.
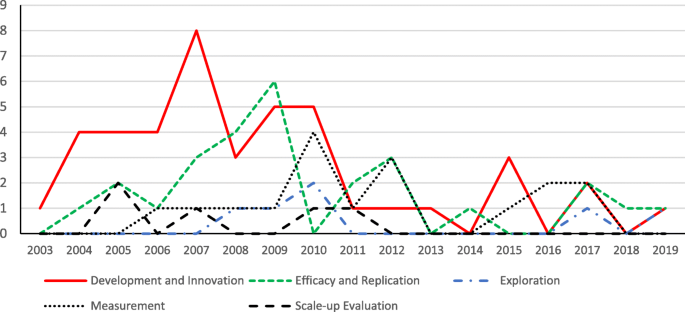
The distribution of projects in terms of the type of goals over the years (in “Education Research” program)
Figure 17 shows the distribution of project types in the category “Special Education Research.” The pattern is different from the pattern shown in Fig. 16 . The types of “development and innovation” and “efficacy and replication” projects were also the dominant types of projects under “Special Education Research” program category in most of these years from 2007 to 2019. Projects in the type “measurement” were only observed in 2010 when that was the only type of project funded.
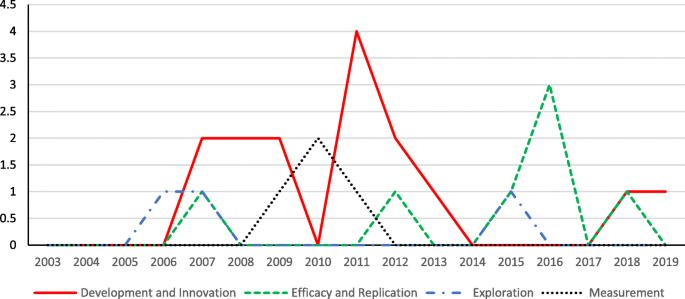
The distribution of projects in terms of goals over the years (in “Special Education Research” program)
Question 6: disciplinary foci of projects in developing and conducting STEM education research
Figure 18 shows that the majority of the 127 projects under such specific programs included disciplinary foci on individual STEM disciplines: mathematics in 88 projects, science in 51 projects, technology in 43 projects, and engineering in 2 projects. The tremendous attention to mathematics in these projects is a bit surprising, as mathematics was noted as being out of balance in STEM education (English, 2016 ) and also in STEM education publications (Li, 2018b , 2019 ). As noted above, each project can be classified in multiple disciplinary foci. However, of the 88 projects with a disciplinary focus on mathematics, 54 projects had mathematics as the only disciplinary focus (38 under “Education Research” program and 16 under “Special Education Research” program). We certainly hope that there will be more projects that further scholarship where mathematics is included as part of (integrated) STEM education (see Li & Schoenfeld, 2019 ).
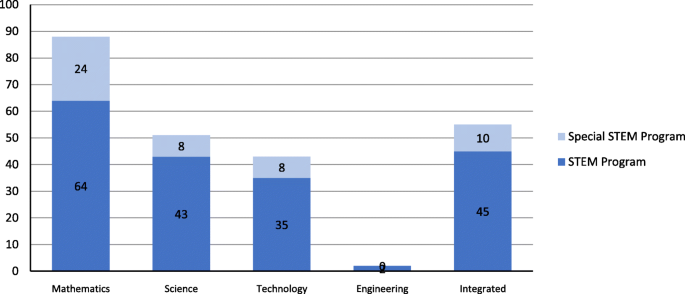
The number of projects in terms of disciplinary focus
There were also projects with specific focus on integrated STEM education (i.e., combining any two or more disciplines of STEM), with a total of 55 (43.3%) projects in a combined program count. The limited number of projects on integrated STEM in the designated STEM funding programs further confirms the common perception that the development of integrated STEM education and research is still in its initial stage (Honey et al., 2014 ; Li, 2018a ).
In examining possible funding trends, Fig. 19 shows that mathematics projects were more frequently funded before 2012. Engineering was a rare disciplinary focus. Integrated STEM was a disciplinary focus from time to time among these projects. No other trends were observed.
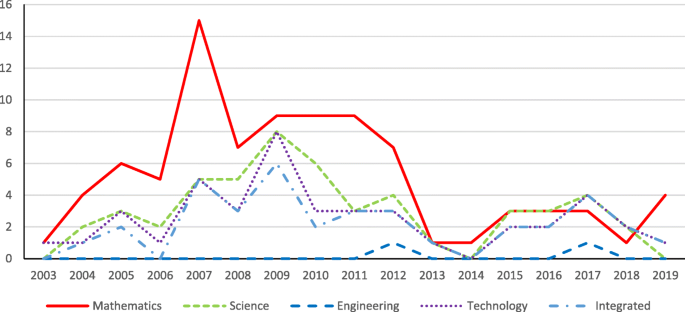
The distribution of projects in terms of disciplinary focus over the years
Question 7: research types and methods that projects used
Figure 20 indicates that “interventional” (in 104 projects, 81.9%) and “experimental research” (in 89 projects, 70.1%) were the most frequently funded types of research. The percentages of projects funded under the regular education research program were similar to those funded under “Special Education Research” program, except that projects funded under “Special Education Research” tended to utilize correlational research more often.
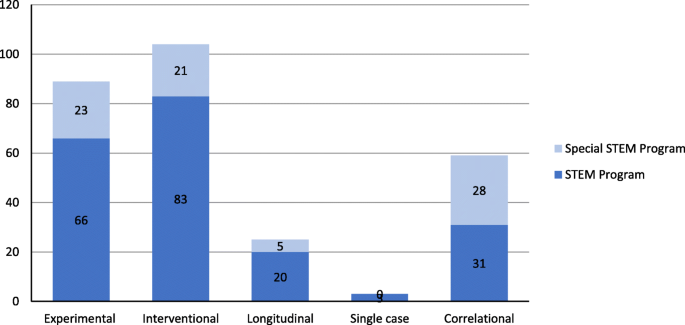
The number of projects in terms of the type of research conducted
Research in STEM education uses diverse data collection and analysis methods; therefore, we wanted to study types of methods (Figs. 21 and 22 , respectively). Among the six types of methods used for data collection, Fig. 21 indicates that “standardized tests” and “designed tests” were the most commonly used methods for data collection, followed by “survey,” “observation,” and “interview.” The majority of projects used three quantitative methods (“standardized tests,” “researcher designed tests,” and “survey”). The finding is consistent with the finding from analysis of journal publications in STEM education (Li et al., 2020 ). Data collected through “interview” and “observation” were more likely to be analyzed using qualitative methods as part of a project’s research methodology.
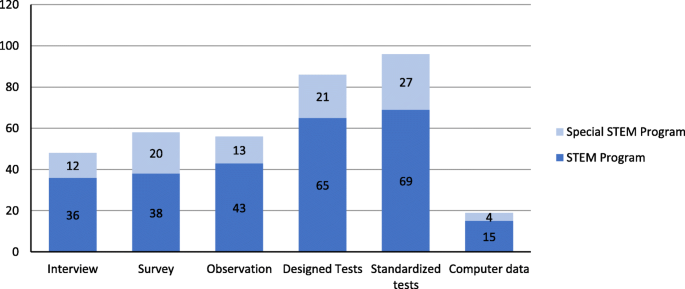
The number of projects categorized by the type of data collection methods

The number of projects categorized by the type of data analysis methods
Figure 22 shows the use of seven (including others) data analysis methods among these projects. The first six methods (i.e., descriptive, ANOVA*, general regression, HLM, IRT, and SEM) as well as some methods in “others” are quantitative data analysis methods. The number of projects that used these quantitative methods is considerably larger than the number of projects that used qualitative methods (i.e., included in “others” category).
Concluding remarks
The systematic analysis of IES-funded research projects in STEM education presented an informative picture about research support for STEM education development in the USA, albeit based on only one public funding agency from 2003 to 2019. Over this 17-year span, IES funded 127 STEM education research projects (an average of over seven projects per year) in two designated STEM program categories. Although we found no discernable longitudinal funding patterns in these two program categories, both the number of funded projects in STEM education and their funding amounts were high. If we included an additional 52 projects with the acronym “STEM” funded by many other programs from 2007 to 2019 (see “ Notes ” section below), the total number of projects in STEM education research would be even higher, and the number of projects with the acronym “STEM” would also be larger. The results suggested the involvement of many researchers with diverse expertise in STEM education research was supported by a broad array of program areas in IES.
Addressing the seven questions showed several findings. Funding support for STEM education research was strong, with an average of about 2 million USD per project for a typical 3–4 year duration. Also, our analysis showed that the number of projects with multiple PIs over the years increased over the study time period, which we speculate was because STEM education research increasingly requires collaboration. STEM education research is still in early development stage, evidenced by the predominance of project goals in either “development and innovation” or “efficacy and replication” categories. We found very few projects (5 out of 127 projects, 4.0%) that were funded for “scale-up evaluation.” Finally, as shown by our analysis of project participants, IES had focused on funding projects for students in grades 1–12. Various quantitative research methods were frequently used by these projects for data collection and analyses.
These results illustrated how well STEM education research was supported through both the designated STEM education and many other programs during the study time period, which helps to explain why researchers in the USA have been so productive in producing and promoting scholarship in STEM education (Li et al., 2019 ; Li et al., 2020 ). We connected several findings from this study to findings from recent reviews of journal publications in STEM education. For example, publications in STEM education appeared in many different journals as many researchers with diverse expertise were supported to study various issues related to STEM education, STEM education publications often have co-authorship, and there is heavy use of quantitative research methods. The link between public funding and significant numbers of publications in STEM education research from US scholars offers a strong argument for the importance of providing strong funding support to research and development in STEM education in the USA and also in many other countries around the world.
The systematic analysis also revealed that STEM education, as used by IES in naming the designated programs, did not convey a clear definition or scope. In fact, we found diverse disciplinary foci in these projects. Integrated STEM was not a main focus of these designated programs in funding STEM education. Instead, many projects in these programs had clear subject content focus in individual disciplines, which is very similar to discipline-based education research (DBER, National Research Council, 2012 ). Interestingly enough, STEM education research had also been supported in many other programs of IES with diverse foci Footnote 7 , such as “Small Business Innovation Research,” “Cognition and Student Learning,” and “Postsecondary and Adult Education.” This funding reality further suggested the broad scope of issues associated with STEM education, as well as the growing need of building STEM education research as a distinct field (Li, 2018a ).
Inspired by our recent review of journal publications as research output in STEM education, this review started with an ambitious goal to study funding support as research input for STEM education. However, we had to limit the scope of the study for feasibility. We limited funding sources to one federal agency in the USA. Therefore, we did not analyze funding support from private funding sources including many private foundations and corporations. Although public funding sources have been one of the most important funding supports available for researchers to develop and expand their research work, the results of this systematic analysis suggest the importance future studies to learn more about research support and input to STEM education from other sources including other major public funding agencies, private foundations, and non-profit professional organizations.
Among these 54 funded projects containing the acronym “STEM” from 2007 to 2019, Table 1 shows that only 2 (3.7%) were in the IES designated program option of STEM education in the category of “Education Research.” Forty-nine projects were in 13 other program options in the category of “Education Research,” with surprisingly large numbers of projects under the “Small Business Innovation Research” option (17, 31.5%) and “Cognition and Student Learning” (11, 20.4%). Three of the 54 funded projects were in the program category of “Special Education Research.” To be specific, two of the three were in the program of “Small Business Innovation Research in Special Education,” and one was in the program of “Special Topic: Career and Technical Education for Students with Disabilities.”
The results suggest that many projects, focusing on various issues and questions directly associated with STEM education, were funded even when researchers applied for funding support in program options not designated as “Science, Technology, Engineering, and Mathematics (STEM) Education.” It implies that issues associated with STEM education had been generally acknowledged as important across many different program areas in education research and special education research. The funding support available in diverse program areas likely allowed numerous scholars with diverse expertise to study many different questions and publish their research in diverse journals, as we noted in the recent review of journal publications in STEM education (Li et al., 2020 ).
A previous study identified and analyzed a total of 46 IES funded projects from 2007 to 2018 (with an average of fewer than 4 projects per year) that contain the acronym “STEM” in a project’s title and/or description (Wang et al., 2019 ). Finding eight newly funded projects in 2019 suggested a growing interest in research on issues directly associated with STEM education in diverse program areas. In fact, five out of these eight newly funded projects specifically included the acronym “STEM” in the project’s title to explicitly indicate the project’s association with STEM education.
Availability of data and materials
The data and materials used and analyzed for the review are publicly available at the IES website, White House website, and other government agency websites.
In a previous study (Wang, Li, & Xiao, 2019), we used the acronym “STEM” as a search term under the option of “SEARCH FUNDED RESEARCH GRANTS AND CONTRACTS” without any program category restriction, and identified and analyzed 46 funded projects from 2007 to 2018 that contain “STEM” in a project’s title and/or description after screening out unrelated key words containing “stem” such as “system”. To make comparisons when needed, we did the same search using the acronym “STEM” and found 8 more funded projects in 2019 for a total of 54 funded projects across many different program categories from 2007 to 2019.
The project of “A Randomized Controlled Study of the Effects of Intelligent Online Chemistry Tutors in Urban California School Districts” (2008). In the project description, its subtitle shows intervention information. We coded this project as “interventional.” Then, the project also included the treatment group and the control group. We coded this project as “experimental.” Finally, this project was to test the efficacy of computer-based cognitive tutors. This was a correlational study. We thus coded it as “correlational.”
Computer data means that the project description indicated this kind of information, such as log data on students.
Descriptive means “descriptive statistics.” General regression means multiple regression, linear regression, logistical regression, except hierarchical linear regression model. ANOVA* is used here as a broad term to include analysis of variance, analysis of covariance, multivariate analysis of variance, and/or multivariate analysis of variance. Others include factor analysis, t tests, Mann-Whitney tests, and binomial tests, log data analysis, meta-analysis, constant comparative data analysis, and qualitative analysis.
Special education originally was about students with disabilities. It has broadened in scope over the years.
The number of students under Special Education was 14% of students in public schools in the USA in 2017–2018. https://nces.ed.gov/programs/coe/indicator_cgg.asp
For example, “Design Environment for Educator-Student Collaboration Allowing Real-Time Engineering-centric, STEM (DESCARTES) Exploration in Middle Grades” (2017) was funded as a 2-year project to Parametric Studios, Inc. (awardee) under the program option of “Small Business Innovation Research” (here is the link: https://ies.ed.gov/funding/grantsearch/details.asp?ID=1922 ). “Exploring the Spatial Alignment Hypothesis in STEM Learning Environments” (2017) was funded as a 4-year project to WestEd (awardee) under the program option of “Cognition and Student Learning” (link: https://ies.ed.gov/funding/grantsearch/details.asp?ID=2059 ). “Enhancing Undergraduate STEM Education by Integrating Mobile Learning Technologies with Natural Language Processing” (2018) was funded as a 4-year project to Purdue University (awardee) under the program option of “Postsecondary and Adult Education” (link: https://ies.ed.gov/funding/grantsearch/details.asp?ID=2130 ).
Abbreviations
Analysis of variance
Discipline-based education research
Department of Education
Hierarchical linear modeling
Institute of Education Sciences
Item response theory
National Science Foundation
Pre-school–grade 12
Requests-for-proposal
Structural equation modeling
Science, technology, engineering, and mathematics
Committee on STEM Education, National Science & Technology Council, the White House (2018). Charting a course for success: America’s strategy for STEM education . Washington, DC. https://www.whitehouse.gov/wp-content/uploads/2018/12/STEM-Education-Strategic-Plan-2018.pdf Accessed on 18 Jan 2019.
English, L. D. (2016). STEM education K-12: perspectives on integration. International Journal of STEM Education, 3 , 3 https://doi.org/10.1186/s40594-016-0036-1 .
Article Google Scholar
Honey, M., Pearson, G., & Schweingruber, A. (2014). STEM integration in K-12 education: status, prospects, and an agenda for research . Washington DC: National Academies Press.
Google Scholar
Institute of Education Sciences (IES) (n.d.). About IES: connecting research, policy and practice. Retrieved from https://ies.ed.gov/aboutus/ Accessed on 2 Feb 2020.
Institute of Education Sciences (IES) & National Science Foundation (NSF). (2013). Common guidelines for education research and development. Washington, DC: The authors. Retrieved from https://www.nsf.gov/pubs/2013/nsf13126/nsf13126.pdf Accessed on 2 Feb 2020.
Lee, A. (2014). Students with disabilities choosing science technology engineering and math (STEM) majors in postsecondary institutions. Journal of Postsecondary Education and Disability, 27 (3), 261–272.
Li, Y. (2014). International journal of STEM education – a platform to promote STEM education and research worldwide. International Journal of STEM Education, 1 , 1 https://doi.org/10.1186/2196-7822-1-1 .
Li, Y. (2018a). Journal for STEM Education Research – promoting the development of interdisciplinary research in STEM education. Journal for STEM Education Research, 1 (1-2), 1–6 https://doi.org/10.1007/s41979-018-0009-z .
Li, Y. (2018b). Four years of development as a gathering place for international researchers and readers in STEM education. International Journal of STEM Education, 5 , 54 https://doi.org/10.1186/s40594-018-0153-0 .
Li, Y. (2019). Five years of development in pursuing excellence in quality and global impact to become the first journal in STEM education covered in SSCI. International Journal of STEM Education, 6 , 42 https://doi.org/10.1186/s40594-019-0198-8 .
Li, Y., Froyd, J. E., & Wang, K. (2019). Learning about research and readership development in STEM education: a systematic analysis of the journal’s publications from 2014 to 2018. International Journal of STEM Education, 6 , 19 https://doi.org/10.1186/s40594-019-0176-1 .
Li, Y., & Schoenfeld, A. H. (2019). Problematizing teaching and learning mathematics as ‘given’ in STEM education. International Journal of STEM Education, 6 , 44 https://doi.org/10.1186/s40594-019-0197-9 .
Li, Y., Wang, K., Xiao, Y., & Froyd, J. E. (2020). Research and trends in STEM education: a systematic review of journal publications. International Journal of STEM Education, 7 , 11 https://doi.org/10.1186/s40594-020-00207-6 .
Margot, K. C., & Kettler, T. (2019). Teachers’ perception of STEM integration and education: a systematic literature review. International Journal of STEM Education, 6 , 2 https://doi.org/10.1186/s40594-018-0151-2 .
Minichiello, A., Hood, J. R., & Harkness, D. S. (2018). Bring user experience design to bear on STEM education: a narrative literature review. Journal for STEM Education Research, 1 (1-2), 7–33.
National Research Council. (2012). Discipline-based education research: understanding and improving learning in undergraduate science and engineering . Washington DC: National Academies Press.
Otten, M., Van den Heuvel-Panhuizen, M., & Veldhuis, M. (2019). The balance model for teaching linear equations: a systematic literature review. International Journal of STEM Education, 6 , 30 https://doi.org/10.1186/s40594-019-0183-2 .
Schreffler, J., Vasquez III, E., Chini, J., & James, W. (2019). Universal design for learning in postsecondary STEM education for students with disabilities: a systematic literature review. International Journal of STEM Education, 6 , 8 https://doi.org/10.1186/s40594-019-0161-8 .
The White House (2009). President Obama launches “Educate to Innovate” campaign for excellence in science, technology, engineering & math (Stem) education. Retrieved from https://obamawhitehouse.archives.gov/the-press-office/president-obama-launches-educate-innovate-campaign-excellence-science-technology-en Accessed on 2 Feb 2020.
The White House (2017). Presidential memorandum for the secretary of Education. Retrieved from https://www.whitehouse.gov/presidential-actions/presidential-memorandum-secretary-education/ Accessed on 2 Feb 2020.
The White House (2018). President Donald J. Trump is working to ensure all Americans have access to STEM education. Retrieved from https://www.whitehouse.gov/briefings-statements/president-donald-j-trump-is-working-to-ensure-all-americans-have-access-to-stem-education/ Accessed on 2 Feb 2020.
U.S. Department of Education (2018). U.S. Department of Education fulfills administration promise to invest $200 million in STEM education. Retrieved from https://www.ed.gov/news/press-releases/us-department-education-fulfills-administration-promise-invest-200-million-stem-education Accessed on 2 Feb 2020.
Wang, K., Li, Y., & Xiao, Y. (2019). Exploring the status and development trends of STEM education research: the case of IES funded projects on STEM education in the U.S. 数学教育学报 . Journal of Mathematics Education, 28 (3), 53–61.
Download references
This review was supported by a grant from the National Science Foundation (DUE-1852942). Any opinions, findings, and conclusions or recommendations expressed in this material are those of the author(s) and do not necessarily reflect the views of the National Science Foundation.
Author information
Authors and affiliations.
Texas A&M University, College Station, TX, 77843-4232, USA
Yeping Li, Yu Xiao & Sandra B. Nite
Nicholls State University, Thibodaux, LA, 70310, USA
Ohio State University, Columbus, OH, 43210, USA
Jeffrey E. Froyd
You can also search for this author in PubMed Google Scholar
Contributions
YL conceptualized the study and drafted the manuscript. KW contributed with data collection, coding, analyses, and manuscript reviews. YX contributed to data collection, coding, and manuscript reviews. JEF and SBN contributed to manuscript improvement through manuscript reviews and revisions. All authors read and approved the final manuscript.
Corresponding author
Correspondence to Yeping Li .
Ethics declarations
Competing interests.
The authors declare that they have no competing interests.
Additional information
Publisher’s note.
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/ .
Reprints and permissions
About this article
Cite this article.
Li, Y., Wang, K., Xiao, Y. et al. Research and trends in STEM education: a systematic analysis of publicly funded projects. IJ STEM Ed 7 , 17 (2020). https://doi.org/10.1186/s40594-020-00213-8
Download citation
Received : 18 March 2020
Accepted : 20 March 2020
Published : 22 April 2020
DOI : https://doi.org/10.1186/s40594-020-00213-8
Share this article
Anyone you share the following link with will be able to read this content:
Sorry, a shareable link is not currently available for this article.
Provided by the Springer Nature SharedIt content-sharing initiative
- Scholarship
- STEM education research
- STEM funding
Advertisement
Trends and Hot Topics of STEM and STEM Education: a Co-word Analysis of Literature Published in 2011–2020
- Published: 23 February 2023
- Volume 33 , pages 1069–1092, ( 2024 )
Cite this article

- Ying-Shao Hsu ORCID: orcid.org/0000-0002-1635-8213 1 , 2 ,
- Kai-Yu Tang ORCID: orcid.org/0000-0002-3965-3055 3 &
- Tzu-Chiang Lin ORCID: orcid.org/0000-0003-3842-3749 4 , 5
1223 Accesses
3 Citations
Explore all metrics
This study explored research trends in science, technology, engineering, and mathematics (STEM) education. Descriptive analysis and co-word analysis were used to examine articles published in Social Science Citation Index journals from 2011 to 2020. From a search of the Web of Science database, a total of 761 articles were selected as target samples for analysis. A growing number of STEM-related publications were published after 2016. The most frequently used keywords in these sample papers were also identified. Further analysis identified the leading journals and most represented countries among the target articles. A series of co-word analyses were conducted to reveal word co-occurrence according to the title, keywords, and abstract. Gender moderated engagement in STEM learning and career selection. Higher education was critical in training a STEM workforce to satisfy societal requirements for STEM roles. Our findings indicated that the attention of STEM education researchers has shifted to the professional development of teachers. Discussions and potential research directions in the field are included.
This is a preview of subscription content, log in via an institution to check access.
Access this article
Subscribe and save.
- Get 10 units per month
- Download Article/Chapter or eBook
- 1 Unit = 1 Article or 1 Chapter
- Cancel anytime
Price includes VAT (Russian Federation)
Instant access to the full article PDF.
Rent this article via DeepDyve
Institutional subscriptions
Similar content being viewed by others

An Integrative Review with Word Cloud Analysis of STEM Education
A review of STEM education with the support of visualizing its structure through the CiteSpace software
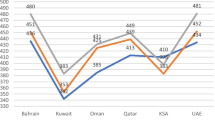
A systematic review of STEM education research in the GCC countries: trends, gaps and barriers
Data availability.
The datasets generated during and/or analysed during the current study are available from the corresponding author on reasonable request.
Akgunduz, D. (2016). A Research about the placement of the top thousand students placed in STEM fields in Turkey between the years 2000 and 2014. EURASIA Journal of Mathematics, Science and Technology Education, 12 (5), 1365–1377.
Google Scholar
Appianing, J., & Van Eck, R. N. (2018). Development and validation of the Value-Expectancy STEM Assessment Scale for students in higher education. International Journal of STEM Education , 5 , article 24.
Assefa, S. G., & Rorissa, A. (2013). A bibliometric mapping of the structure of STEM education using co-word analysis. Journal of the American Society for Information Science and Technology, 64 (12), 2513–2536.
Belland, B. R., Walker, A. E., Kim, N. J., & Lefler, M. (2017). Synthesizing results from empirical research on computer-based scaffolding in STEM education: A meta-analysis. Review of Educational Research, 87 (2), 309–344.
Brotman, J. S., & Moore, F. M. (2008). Girls and science: A review of four themes in the science education literature. Journal of Research in Science Teaching, 45 (9), 971–1002.
Brown, R. E., & Bogiages, C. A. (2019). Professional development through STEM integration: How early career math and science teachers respond to experiencing integrated STEM tasks. International Journal of Science and Mathematics Education, 17 (1), 111–128.
Burt, B. A., Williams, K. L., & Palmer, G. J. M. (2019). It takes a village: The role of emic and etic adaptive strengths in the persistence of black men in engineering graduate programs. American Educational Research Journal, 56 (1), 39–74.
Callon, M., Courtial, J. P., & Laville, F. (1991). Co-word analysis as a tool for describing the network of interactions between basic and technological research: The case of polymer chemistry. Scientometrics, 22 (1), 155–205.
Carlisle, D. L. & Weaver, G. C. (2018). STEM education centers: Catalyzing the improvement of undergraduate STEM education. International Journal of STEM Education, 5 , article 47.
Chang, D. F., & ChangTzeng, H. C. (2020). Patterns of gender parity in the humanities and STEM programs: The trajectory under the expanded higher education system. Studies in Higher Education, 45 (6), 1108–1120.
Charleston, L. J. (2012). A qualitative investigation of African Americans’ decision to pursue computing science degrees: Implications for cultivating career choice and aspiration. Journal of Diversity in Higher Education, 5 (4), 222–243.
Charleston, L. J., George, P. L., Jackson, J. F. L., Berhanu, J., & Amechi, M. H. (2014). Navigating underrepresented STEM spaces: Experiences of black women in US computing science higher education programs who actualize success. Journal of Diversity in Higher Education, 7 (3), 166–176.
Chien, Y. H., & Chu, P. Y. (2018). The different learning outcomes of high school and college students on a 3D-printing STEAM engineering design curriculum. International Journal of Science and Mathematics Education, 16 (6), 1047–1064.
Dehdarirad, T., Villarroya, A., & Barrios, M. (2014). Research trends in gender differences in higher education and science: A co-word analysis. Scientometrics, 101 (1), 273–290.
Dickerson, D. L., Eckhoff, A., Stewart, C. O., Chappell, S., & Hathcock, S. (2014). The examination of a pullout STEM program for urban upper elementary students. Research in Science Education, 44 (3), 483–506.
Eccles, J., Adler, T. F., Futterman, R., Goff, S. B., Kaczala, C. M., Meece, J., & Midgley, C. (1983). Expectancies, values and academic behaviors. In J. T. Spence (Ed.), Achievement and Achievement Motives . W. San Francisco: H. Freeman.
Ellison, S., & Allen, B. (2018). Disruptive innovation, labor markets, and Big Valley STEM School: Network analysis in STEM education. Cultural Studies of Science Education, 13 (1), 267–298.
Erdogan, N., Navruz, B., Younes, R., & Capraro, R. M. (2016). Viewing how STEM project-based learning influences students’ science achievement through the implementation lens: A latent growth modeling. Eurasia Journal of Mathematics, Science and Technology Education, 12 (8), 2139–2154.
European Commission, Directorate-General for Education, Youth, Sport and Culture (2016). Does the EU need more STEM graduates? Final report . Retrieve from https://data.europa.eu/doi/10.2766/000444
Fredricks, J. A., Hofkens, T., Wang, M. T., Mortenson, E., & Scott, P. (2018). Supporting girls’ and boys’ engagement in math and science learning: A mixed methods study. Journal of Research in Science Teaching, 55 (2), 271–298.
Fry, R., Kennedy, B., & Funk, C. (2021). Stem jobs see uneven progress in increasing gender, racial and ethnic diversity. Retrieve from https://www.pewresearch.org/science/wp-content/uploads/sites/16/2021/03/PS_2021.04.01_diversity-in-STEM_REPORT.pdf
Ganley, C. M., George, C. E., Cimpian, J. R., & Makowski, M. B. (2018). Gender equity in college majors: Looking beyond the STEM/non-STEM dichotomy for answers regarding female participation. American Educational Research Journal, 55 (3), 453–487.
Gehrke, S., & Kezar, A. (2019). Perceived outcomes associated with engagement in and design of faculty communities of practice focused on STEM reform. Research in Higher Education, 60 (4), 844–869.
Gilmore, J., Vieyra, M., Timmerman, B., Feldon, D., & Maher, M. (2015). The relationship between undergraduate research participation and subsequent research performance of early career STEM graduate students. Journal of Higher Education, 86 (6), 834–863.
Godwin, A., Potvin, G., Hazari, Z., & Lock, R. (2016). Identity, critical agency, and engineering: An affective model for predicting engineering as a career choice. Journal of Engineering Education, 105 (2), 312–340.
Han, S., Yalvac, B., Capraro, M. M., & Capraro, R. M. (2015). In-service teachers’ implementation and understanding of STEM project based learning. Eurasia Journal of Mathematics Science and Technology Education, 11 (1), 63–76.
Heras, M., Ruiz-Mallén, I., & Gallois, S. (2020). Staging science with young people: Bringing science closer to students through stand-up comedy. International Journal of Science Education, 42 (12), 1968–1987.
Hernandez, P. R., Estrada, M., Woodcock, A., & Schultz, P. W. (2017). Protégé perceptions of high mentorship quality depend on shared values more than on demographic match. Journal of Experimental Education, 85 (3), 450–468.
Hinojo Lucena, F. J., Lopez Belmonte, J., Fuentes Cabrera, A., Trujillo Torres, J. M., & Pozo Sanchez, S. (2020). Academic effects of the use of flipped learning in physical education. International journal of Environmental Research and Public Health , 17 (1), article 276.
Holmes, K., Gore, J., Smith, M., & Lloyd, A. (2018). An integrated analysis of school students’ aspirations for STEM careers: Which student and school factors are most predictive? International Journal of Science and Mathematics Education, 16 (4), 655–675.
Huang, X., & Qiao, C. (2022). Enhancing computational thinking skills through artificial intelligence education at a STEAM high school. Science & Education . https://doi.org/10.1007/s11191-022-00392-6
Article Google Scholar
Hughes, R. M., Nzekwe, B., & Molynearx, K. J. (2013). The single sex debate for girls in science: A comparison between two informal science programs on middle school students’ STEM identity formation. Research in Science Education, 43 , 1979–2007.
Hughes, B. S., Corrigan, M. W., Grove, D., Andersen, S. B., & Wong, J. T. (2022). Integrating arts with STEM and leading with STEAM to increase science learning with equity for emerging bilingual learners in the United States. International Journal of STEM Education , 9 , article 58.
Johnson, A. M. (2019). “I can turn it on when I need to”: Pre-college integration, culture, and peer academic engagement among black and Latino/a engineering students. Sociology of Education, 92 (1), 1–20.
Kayan-Fadlelmula, F., Sellami, A., Abdelkader, N., & Umer, S. (2022). A systematic review of STEM education research in the GCC countries: Trends, gaps and barriers. International Journal of STEM Education, 9 , article 2.
Kelly, R., Mc Garr, O., Leahy, K., & Goos, M. (2020). An investigation of university students and professionals’ professional STEM identity status. Journal of Science Education and Technology, 29 (4), 536–546.
Kezar, A., Gehrke, S., & Bernstein-Sierra, S. (2017). Designing for success in STEM communities of practice: Philosophy and personal interactions. The Review of Higher Education, 40 (2), 217–244.
Kezar, A., Gehrke, S., & Bernstein-Sierra, S. (2018). Communities of transformation: Creating changes to deeply entrenched issues. The Journal of Higher Education, 89 (6), 832–864.
Kricorian, K., Seu, M., Lpoez, D., Ureta, E., & Equils, O. (2020). Factors influencing participation of underrepresented students in STEM fields: Matched mentors and mindsets. International Journal of STEM Education, 7 , article 16.
Ku, C. J., Hsu, Y. S., Chang, M. C., & Lin, K. Y. (2022). A model for examining middle school students’ STEM integration behavior in a national technology competition. International Journal of STEM Education, 9 (1), 3.
Leydesdroff, L. (1989). Words and co-words as indicators of intellectual organization. Research Policy, 18 (4), 209–223.
Li, Y., Wang, K., Xiao, Y., & Froyd, J. E. (2020a). Research and trends in STEM education: A systematic review of journal publications. International Journal of STEM Education, 7 , article 11.
Li, Y., Wang, K., Xiao, Y., Froyd, J. E., Nite, S. B. (2020b). Research and trends in STEM education: A systematic analysis of publicly funded projects. International Journal of STEM Education, 7 , article 17.
Lin, T. C., Lin, T. J., & Tsai, C. C. (2014). Research trends in science education from 2008 to 2012: A systematic content analysis of publications in selected journals. International Journal of Science Education, 36 (8), 1346–1372.
Lin, T. J., Lin, T. C., Potvin, P., & Tsai, C. C. (2019). Research trends in science education from 2013 to 2017: A systematic content analysis of publications in selected journals. International Journal of Science Education, 41 (3), 367–387.
Lin, T. C., Tang, K. Y., Lin, S. S., Changlai, M. L., & Hsu, Y. S. (2022). A co-word analysis of selected science education literature: Identifying research trends of scaffolding in two decades (2000–2019). Frontiers in Psychology, 13 , 844425.
Liu, J. S., & Lu, L. Y. (2012). An integrated approach for main path analysis: Development of the Hirsch index as an example. Journal of the American Society for Information Science and Technology, 63 (3), 528–542.
Liu, C. Y., & Wu, C. J. (2022). STEM without art: A ship without a sail. Thinking Skills and Creativity, 43 , 100977.
Lou, S. H., Shih, R. C., Diez, C. R., & Tseng, K. H. (2011). The impact of problem-based learning strategies on STEM knowledge integration and attitudes: An exploratory study among female Taiwanese senior high school students. International Journal of Technology and Design Education, 21 (2), 195–215.
Lynch, S. J., Burton, E. P., Behrend, T., House, A., Ford, M., Spillane, N., Matray, S., Han, E., & Means, B. (2018). Understanding inclusive STEM high schools as opportunity structures for underrepresented students: Critical components. Journal of Research in Science Teaching, 55 (5), 712–748.
Maass, K., Geiger, V., Ariza, M. R., & Goos, M. (2019). The Role of mathematics in interdisciplinary STEM education. ZDM-Mathematics Education, 51 (6), 869–884.
Mansfield, K. C. (2014). How listening to student voices informs and strengthens social justice research and practice. Educational Administration Quarterly, 50 (3), 392–430.
Margot, K. C., & Kettler, T. (2019). Teachers’ perception of STEM integration and education: A systematic literature review. International Journal of STEM education , 6 , article 2.
Marín-Marín, J. A., Moreno-Guerrero, A. J., Dúo-Terrón, P., & López-Belmonte, J. (2021). STEAM in education: A bibliometric analysis of performance and co-words in Web of Science. International Journal of STEM Education , 8 , article 41.
Martín-Páez, T., Aguilera, D., Perales-Palacios, F. J., & Vílchez-González, J. M. (2019). What are we talking about when we talk about STEM education? A Review of Literature. Science Education, 103 (4), 799–822.
McGee, E. O. (2020). Interrogating structural racism in STEM higher education. Educational Researcher, 49 (9), 633–644.
Meho, L. I., & Yang, K. (2006). A new era in citation and bibliometric analyses: Web of Science, Scopus, and Google Scholar. arXiv preprint cs/0612132 .
Mejias, S., Thompson, N., Sedas, R. M., Rosin, M., Soep, E., Peppler, K., Roche, J., Wong, J., Hurley, M., Bell, P., & Bevan, B. (2021). The trouble with STEAM and why we use it anyway. Science Education, 105 (2), 209–231.
Micari, M., Van Winkle, Z., & Pazos, P. (2016). Among friends: The role of academic-preparedness diversity in individual performance within a small-group STEM learning environment. International Journal of Science Education, 38 (12), 1904–1922.
Millar, V. (2020). Trends, issues and possibilities for an interdisciplinary STEM curriculum. Science & Education, 29 (4), 929–948.
Nadelson, L. S., Callahan, J., Pyke, P., Hay, A., Dance, M., & Pfiester, J. (2013). Teacher STEM perception and preparation: Inquiry-based STEM professional development for elementary teachers. Journal of Educational Research, 106 (2), 157–168.
Nakatoh, T., & Hirokawa, S. (2019, July). Evaluation index to find relevant papers: Improvement of focused citation count. In International Conference on Human-Computer Interaction (pp. 555–566). Springer, Cham.
National Science Technology Council. (2012). Coordinating federal science, technology, engineering, and mathematics (STEM) education investments: Progress report. Retrieved from https://obamawhitehouse.archives.gov/sites/default/files/microsites/ostp/nstc_federal_stem_education_coordination_report.pdf
National Science Technology Council. (2013). Federal Science, Technology, Engineering, and Mathematics (STEM) Education 5-Year Strategic Plan. Retrieved from https://obamawhitehouse.archives.gov/sites/default/files/microsites/ostp/stem_stratplan_2013.pdf
Ong, M., Smith, J. M., & Ko, L. T. (2018). Counterspaces for women of color in STEM higher education: Marginal and central spaces for persistence and success. Journal of Research in Science Teaching, 55 (2), 206–245.
Organisation for Economic Cooperation and Development, OECD (2021). Education at A Glance 2021. Retrieve from https://read.oecd.org/10.1787/b35a14e5-en?format=pdf
Perez-Felkner, L., Felkner, J. S., Nix, S., & Magalhaes, M. (2020). The puzzling relationship between international development and gender equity: The case of STEM postsecondary education in Cambodia. International Journal of Educational Development, 72 , 102102.
Perignat, E., & Katz-Buonincontro, J. (2019). STEAM in practice and research: An integrative literature review. Thinking Skills and Creativity, 31 , 31–43.
Quigley, C. F., & Herro, D. (2016). “Finding the joy in the unknown”: Implementation of steam teaching practices in middle school science and math classrooms. Journal of Science Education and Technology, 25 (3), 410–426.
Ramey, K. E., & Stevens, R. (2019). Interest development and learning in choice-based, in-school, making activities: The case of a 3D printer. Learning, Culture and Social Interaction, 23 , 100262.
Salami, M. K., Makela, C. J., & de Miranda, M. A. (2017). Assessing changes in teachers’ attitudes toward interdisciplinary STEM teaching. International Journal of Technology and Design Education, 27 (1), 63–88.
Sanders, M. (2009). Integrative STEM education primer. The Technology Teacher, 68 (4), 20–26.
Saorín, J. L., Melian-Díaz, D., Bonnet, A., Carrera, C. C., Meier, C., & De La Torre-Cantero, J. (2017). Makerspace teaching-learning environment to enhance creative competence in engineering students. Thinking Skills and Creativity, 23 , 188–198.
Simon, R. M., Wagner, A., & Killion, B. (2017). Gender and choosing a STEM major in college: Femininity, masculinity, chilly climate, and occupational values. Journal of Research in Science Teaching, 54 (3), 299–323.
Stolle-McAllister, K., Domingo, M. R. S., & Carrillo, A. (2011). The Meyerhoff way: How the Meyerhoff scholarship program helps black students succeed in the sciences. Journal of Science Education and Technology, 20 (1), 5–16.
Thomas, B., & Watters, J. J. (2015). Perspectives on Australian, Indian and Malaysian approaches to STEM education. International Journal of Educational Development, 45 , 42–53.
Tosun, C. (2022). Analysis of the last 40 years of science education research via bibliometric methods. Science & Education . https://doi.org/10.1007/s11191-022-00400-9
Van Eck, N. J., & Waltman, L. (2010). Software survey: VOSviewer, a computer program for bibliometric mapping. Scientometrics, 84 (2), 523–538.
Vencent-Ruz, P., & Schunn, C. D. (2017). The increasingly important role of science competency beliefs for science learning in girls. Journal of Research in Science Teaching, 54 (6), 790–822.
Wang, S., Chen, Y., Lv, X., & Xu, J. (2022). Hot topics and frontier evolution of science education research: A bibliometric mapping from 2001 to 2020. Science & Education . https://doi.org/10.1007/s11191-022-00337-z
Weeden, K. A., Gelbgiser, D., & Morgan, S. L. (2020). Pipeline dreams: Occupational plans and gender differences in STEM major persistence and completion. Sociology of Education, 93 (4), 297–314.
Wigfield, A., & Eccles, J. S. (2000). Expectancy-value theory of achievement motivation. Contemporary Educational Psychology, 25 (1), 68–81.
Download references
Author information
Authors and affiliations.
Graduate Institute of Science Education, National Taiwan Normal University, No. 88, Ting-Jou Rd., Sec. 4, Taipei City, 116, Taiwan
Ying-Shao Hsu
Institute for Research Excellence in Learning Sciences, National Taiwan Normal University, No. 88, Ting-Jou Rd., Sec. 4, Taipei City, 116, Taiwan
Graduate Institute of Library & Information Science, National Chung Hsing University, 145 Xingda Rd., South Dist., Taichung City, 402, Taiwan
Kai-Yu Tang
Center for Liberal Arts, National Kaohsiung University of Science and Technology, No. 415, Jiangong Rd., Sanmin Dist, Kaohsiung City, 807618, Taiwan
Tzu-Chiang Lin
Center for Teacher Education, National Kaohsiung University of Science and Technology, No. 415, Jiangong Rd., Sanmin Dist, Kaohsiung City, 807618, Taiwan
You can also search for this author in PubMed Google Scholar
Corresponding author
Correspondence to Tzu-Chiang Lin .
Ethics declarations
Ethical approval and consent to participate.
This study involves neither human participants’ data nor relevant biological material. Ethics approval and informed consent are hence not applicable.
Conflict of Interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's note.
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
Reprints and permissions
About this article
Hsu, YS., Tang, KY. & Lin, TC. Trends and Hot Topics of STEM and STEM Education: a Co-word Analysis of Literature Published in 2011–2020. Sci & Educ 33 , 1069–1092 (2024). https://doi.org/10.1007/s11191-023-00419-6
Download citation
Accepted : 26 January 2023
Published : 23 February 2023
Issue Date : August 2024
DOI : https://doi.org/10.1007/s11191-023-00419-6
Share this article
Anyone you share the following link with will be able to read this content:
Sorry, a shareable link is not currently available for this article.
Provided by the Springer Nature SharedIt content-sharing initiative
- STEM education
- Co-word analysis
- Research trends
- Find a journal
- Publish with us
- Track your research
Featured Topics
Featured series.
A series of random questions answered by Harvard experts.
Explore the Gazette
Read the latest.

Toll of QAnon on families of followers

The urgent message coming from boys

Your kid can’t name three branches of government? He’s not alone.

Employment in STEM occupations has grown 79 percent in the past three decades, according to U.S. Bureau of Labor Statistics.
iStock by Getty Images
Growing gap in STEM supply and demand
Brigid O’Rourke
Harvard Correspondent
Experts cite online learning, digital tools as ways to build inclusive and equitable STEM workforce
The evolution and impact of STEM education and its accompanying career opportunities reflect a positive in the fields of science, technology, engineering, and mathematics. But as the need grows for a specialized STEM-focused workforce, it’s becoming clear that not everyone has an equal opportunity.
During the Harvard-sponsored talk, “New Pathways to STEM,” panelists cited a large subset of students who are not being fully prepared for STEM careers. They then discussed ways the gap could be closed, pointing to online learning and the rapid advancement of new digital tools as ways to make STEM education more readily available. These new ways of learning, they said, can ultimately expand access to STEM education and create a more inclusive and equitable STEM workforce.
The need for a vast, talented workforce in STEM-related fields has never been more necessary, said Bridget Long, dean of the Harvard Graduate School of Education. Long cited the U.S. Bureau of Labor Statistics, which shows employment in STEM occupations has grown 79 percent in the past three decades. In addition, STEM jobs are projected to grow an additional 11 percent from 2020 to 2030. In Massachusetts alone, “40 percent of all employment revolves around innovation industries, such as clean energy, information technology, defense and advanced manufacturing,” said Long.
But, she added, “the importance of STEM education is about so much more than just jobs. STEM fields demand curious individuals eager to solve the world’s most pressing problems.”
“We need to have a new vision of how we prepare students to think critically about the world … as well as educating a society such that it has scientific literacy,” said Joseph L. Graves Jr., (upper left). Joining Graves were Brigid Long, Mike Edmonson, Amanda Dillingham, and Martin West.
The study of STEM subjects, she continued, teaches critical-thinking skills, and instills a mindset that will help students find success across numerous areas and disciplines. However, Long said, “too often the opportunity to learn and to be inspired by STEM is not available.
“Only 20 percent of high school graduates are prepared for college-level coursework in STEM majors,” she cited, adding, “fewer than half of high schools in the United States even offer computer science classes. So that begs the question — are kids going to be ready to meet the evolving and growing landscape of STEM professions?”
While STEM education opportunities are often scarce for high school students across the board, it’s even more pervasive when you consider how inequitably access is distributed by income, race, ethnicity, or gender. For example, Long said, “Native American, Black and Latinx students are the least likely to attend schools that teach computer science, as are students from rural areas, and [those with] economically disadvantaged backgrounds.
“It’s not surprising that these differences in educational opportunities lead to very large differences in what we see in the labor force. We are shutting students out of opportunity,” she said.
So what can be done to ensure more students from all backgrounds are exposed to a wide variety of opportunities? According to Graduate School of Education Academic Dean Martin West, who is also a member of the Massachusetts Board of Elementary and Secondary Education, a concerted effort is being made at the state level to work with — and through — teachers to convey to students the breadth of STEM opportunities and to assure them that “it’s not all sitting in front of a computer, or being in a science lab, but showing them that there are STEM opportunities in a wide range of fields.”
The relatively recent emergence of digital platforms, such as LabXchange, are helping to bridge the gap. LabXchange is a free online learning tool for science education that allows students, educators, scientists, and researchers to collaborate in a virtual community. The initiative was developed by Harvard University’s Faculty of Arts and Sciences and the Amgen Foundation . It offers a library of diverse content, includes a biotechnology learning resource available in 13 languages, and applies science to real-world issues. Teachers and students from across the country and around the world can access the free content and learn from wherever they are.
Many of the panelists also pointed to the need for steady funding in helping to address the inequities.
“Bottom line, if this nation wants to be a competitive leader in STEM, it has to revitalize its vision of what it needs to do, particularly in the public schools where most Black and brown people are, with regard to producing the human and physical infrastructure to teach STEM,” said Joseph L. Graves Jr., professor of biological sciences, North Carolina Agricultural and Technical State University. Graves is also a member of the Faculty Steering Committee, LabXchange’s Racial Diversity, Equity, and Inclusion in Science Education Initiative.
The panel noted how LabXchange is partnering with scholars from several historically Black colleges and universities to develop new digital learning resources on antiracism in education, science, and public health. The content, which will be freely available and translated into Spanish, is being funded by a $1.2 million grant from the Amgen Foundation. Aside from the highly successful LabXchange program, Mike Edmondson, vice president, Global Field Excellence and Commercial, Diversity Inclusion & Belonging at Amgen, noted the Amgen Biotech Experience and the Amgen Scholars program — both of which help to ensure that everyone has the opportunity to engage in science and to see themselves in a STEM career.
We also have to do a better job at helping people understand that that we cannot afford to fall behind in STEM education, Graves argued. “That means it’s going to cost us some money. So, America needs to be willing to pay … to build out STEM education infrastructure, so that we can produce the number of STEM professionals we need going forward,” he said. “We need to have a new vision of how we prepare students to think critically about the world … as well as educating a society such that it has scientific literacy.”
Amanda Dillingham, the program director of science and biology at East Boston High School, is on the front lines of this challenge, and says she believes that supporting teachers is one of the most critical steps that can be taken to address the issue in the immediate future.
When more funding is brought to the table, teachers “are able to coordinate networks … and build biotech labs in our classrooms and build robotics labs in our classrooms …. and are actually able to introduce students to [these fields and these careers] at a very early age,” said Dillingham.
Long and the panel also paid tribute to Rob Lue, the brainchild behind LabXchange, who passed away a year ago.
“Rob challenged science learners, scientists and educators to commit to ending racial inequity,” Long said. “Access was at the core of all of Rob’s many contributions to education at Harvard and beyond. He envisioned a world without barriers and where opportunity was available to anyone, especially in science. In everything that he did, he created an environment in which learners of all ages of diverse backgrounds could come together to imagine, learn, and achieve live exchange. Rob’s free online learning platform for science was his most expansive vision, and one that continues to inspire educators and learners around the world.”
Share this article
You might like.

New book by Nieman Fellow explores pain, frustration in efforts to help loved ones break free of hold of conspiracy theorists

The urgent message of ‘Rebels with a Cause’? When we don’t listen, we all suffer.

Efforts launched to turn around plummeting student scores in U.S. history, civics, amid declining citizen engagement across nation
You want to be boss. You probably won’t be good at it.
Study pinpoints two measures that predict effective managers
Good genes are nice, but joy is better
Harvard study, almost 80 years old, has proved that embracing community helps us live longer, and be happier
200+ Experimental Quantitative Research Topics For STEM Students In 2023

STEM stands for Science, Technology, Engineering, and Math, but these are not the only subjects we learn in school. STEM is like a treasure chest of skills that help students become great problem solvers, ready to tackle the real world’s challenges.
In this blog, we are here to explore the world of Research Topics for STEM Students. We will break down what STEM really means and why it is so important for students. In addition, we will give you the lowdown on how to pick a fascinating research topic. We will explain a list of 200+ Experimental Quantitative Research Topics For STEM Students.
And when it comes to writing a research title, we will guide you step by step. So, stay with us as we unlock the exciting world of STEM research – it is not just about grades; it is about growing smarter, more confident, and happier along the way.
What Is STEM?
Table of Contents
STEM is Science, Technology, Engineering, and Mathematics. It is a way of talking about things like learning, jobs, and activities related to these four important subjects. Science is about understanding the world around us, technology is about using tools and machines to solve problems, engineering is about designing and building things, and mathematics is about numbers and solving problems with them. STEM helps us explore, discover, and create cool stuff that makes our world better and more exciting.
Why STEM Research Is Important?
STEM research is important because it helps us learn new things about the world and solve problems. When scientists, engineers, and mathematicians study these subjects, they can discover cures for diseases, create new technology that makes life easier, and build things that help us live better. It is like a big puzzle where we put together pieces of knowledge to make our world safer, healthier, and more fun.
- STEM research leads to new discoveries and solutions.
- It helps find cures for diseases.
- STEM technology makes life easier.
- Engineers build things that improve our lives.
- Mathematics helps us understand and solve complex problems.
How to Choose a Topic for STEM Research Paper
Here are some steps to choose a topic for STEM Research Paper:
Step 1: Identify Your Interests
Think about what you like and what excites you in science, technology, engineering, or math. It could be something you learned in school, saw in the news, or experienced in your daily life. Choosing a topic you’re passionate about makes the research process more enjoyable.
Step 2: Research Existing Topics
Look up different STEM research areas online, in books, or at your library. See what scientists and experts are studying. This can give you ideas and help you understand what’s already known in your chosen field.
Step 3: Consider Real-World Problems
Think about the problems you see around you. Are there issues in your community or the world that STEM can help solve? Choosing a topic that addresses a real-world problem can make your research impactful.
Step 4: Talk to Teachers and Mentors
Discuss your interests with your teachers, professors, or mentors. They can offer guidance and suggest topics that align with your skills and goals. They may also provide resources and support for your research.
Step 5: Narrow Down Your Topic
Once you have some ideas, narrow them down to a specific research question or project. Make sure it’s not too broad or too narrow. You want a topic that you can explore in depth within the scope of your research paper.
Here we will discuss 200+ Experimental Quantitative Research Topics For STEM Students:
Qualitative Research Topics for STEM Students:
Qualitative research focuses on exploring and understanding phenomena through non-numerical data and subjective experiences. Here are 10 qualitative research topics for STEM students:
- Exploring the experiences of female STEM students in overcoming gender bias in academia.
- Understanding the perceptions of teachers regarding the integration of technology in STEM education.
- Investigating the motivations and challenges of STEM educators in underprivileged schools.
- Exploring the attitudes and beliefs of parents towards STEM education for their children.
- Analyzing the impact of collaborative learning on student engagement in STEM subjects.
- Investigating the experiences of STEM professionals in bridging the gap between academia and industry.
- Understanding the cultural factors influencing STEM career choices among minority students.
- Exploring the role of mentorship in the career development of STEM graduates.
- Analyzing the perceptions of students towards the ethics of emerging STEM technologies like AI and CRISPR.
- Investigating the emotional well-being and stress levels of STEM students during their academic journey.
Easy Experimental Research Topics for STEM Students:
These experimental research topics are relatively straightforward and suitable for STEM students who are new to research:
- Measuring the effect of different light wavelengths on plant growth.
- Investigating the relationship between exercise and heart rate in various age groups.
- Testing the effectiveness of different insulating materials in conserving heat.
- Examining the impact of pH levels on the rate of chemical reactions.
- Studying the behavior of magnets in different temperature conditions.
- Investigating the effect of different concentrations of a substance on bacterial growth.
- Testing the efficiency of various sunscreen brands in blocking UV radiation.
- Measuring the impact of music genres on concentration and productivity.
- Examining the correlation between the angle of a ramp and the speed of a rolling object.
- Investigating the relationship between the number of blades on a wind turbine and energy output.
Research Topics for STEM Students in the Philippines:
These research topics are tailored for STEM students in the Philippines:
- Assessing the impact of climate change on the biodiversity of coral reefs in the Philippines.
- Studying the potential of indigenous plants in the Philippines for medicinal purposes.
- Investigating the feasibility of harnessing renewable energy sources like solar and wind in rural Filipino communities.
- Analyzing the water quality and pollution levels in major rivers and lakes in the Philippines.
- Exploring sustainable agricultural practices for small-scale farmers in the Philippines.
- Assessing the prevalence and impact of dengue fever outbreaks in urban areas of the Philippines.
- Investigating the challenges and opportunities of STEM education in remote Filipino islands.
- Studying the impact of typhoons and natural disasters on infrastructure resilience in the Philippines.
- Analyzing the genetic diversity of endemic species in the Philippine rainforests.
- Assessing the effectiveness of disaster preparedness programs in Philippine communities.
Read More
- Frontend Project Ideas
- Business Intelligence Projects For Beginners
Good Research Topics for STEM Students:
These research topics are considered good because they offer interesting avenues for investigation and learning:
- Developing a low-cost and efficient water purification system for rural communities.
- Investigating the potential use of CRISPR-Cas9 for gene therapy in genetic disorders.
- Studying the applications of blockchain technology in securing medical records.
- Analyzing the impact of 3D printing on customized prosthetics for amputees.
- Exploring the use of artificial intelligence in predicting and preventing forest fires.
- Investigating the effects of microplastic pollution on aquatic ecosystems.
- Analyzing the use of drones in monitoring and managing agricultural crops.
- Studying the potential of quantum computing in solving complex optimization problems.
- Investigating the development of biodegradable materials for sustainable packaging.
- Exploring the ethical implications of gene editing in humans.
Unique Research Topics for STEM Students:
Unique research topics can provide STEM students with the opportunity to explore unconventional and innovative ideas. Here are 10 unique research topics for STEM students:
- Investigating the use of bioluminescent organisms for sustainable lighting solutions.
- Studying the potential of using spider silk proteins for advanced materials in engineering.
- Exploring the application of quantum entanglement for secure communication in the field of cryptography.
- Analyzing the feasibility of harnessing geothermal energy from underwater volcanoes.
- Investigating the use of CRISPR-Cas12 for rapid and cost-effective disease diagnostics.
- Studying the interaction between artificial intelligence and human creativity in art and music generation.
- Exploring the development of edible packaging materials to reduce plastic waste.
- Investigating the impact of microgravity on cellular behavior and tissue regeneration in space.
- Analyzing the potential of using sound waves to detect and combat invasive species in aquatic ecosystems.
- Studying the use of biotechnology in reviving extinct species, such as the woolly mammoth.
Experimental Research Topics for STEM Students in the Philippines
Research topics for STEM students in the Philippines can address specific regional challenges and opportunities. Here are 10 experimental research topics for STEM students in the Philippines:
- Assessing the effectiveness of locally sourced materials for disaster-resilient housing construction in typhoon-prone areas.
- Investigating the utilization of indigenous plants for natural remedies in Filipino traditional medicine.
- Studying the impact of volcanic soil on crop growth and agriculture in volcanic regions of the Philippines.
- Analyzing the water quality and purification methods in remote island communities.
- Exploring the feasibility of using bamboo as a sustainable construction material in the Philippines.
- Investigating the potential of using solar stills for freshwater production in water-scarce regions.
- Studying the effects of climate change on the migration patterns of bird species in the Philippines.
- Analyzing the growth and sustainability of coral reefs in marine protected areas.
- Investigating the utilization of coconut waste for biofuel production.
- Studying the biodiversity and conservation efforts in the Tubbataha Reefs Natural Park.
Capstone Research Topics for STEM Students in the Philippines:
Capstone research projects are often more comprehensive and can address real-world issues. Here are 10 capstone research topics for STEM students in the Philippines:
- Designing a low-cost and sustainable sanitation system for informal settlements in urban Manila.
- Developing a mobile app for monitoring and reporting natural disasters in the Philippines.
- Assessing the impact of climate change on the availability and quality of drinking water in Philippine cities.
- Designing an efficient traffic management system to address congestion in major Filipino cities.
- Analyzing the health implications of air pollution in densely populated urban areas of the Philippines.
- Developing a renewable energy microgrid for off-grid communities in the archipelago.
- Assessing the feasibility of using unmanned aerial vehicles (drones) for agricultural monitoring in rural Philippines.
- Designing a low-cost and sustainable aquaponics system for urban agriculture.
- Investigating the potential of vertical farming to address food security in densely populated urban areas.
- Developing a disaster-resilient housing prototype suitable for typhoon-prone regions.
Experimental Quantitative Research Topics for STEM Students:
Experimental quantitative research involves the collection and analysis of numerical data to conclude. Here are 10 Experimental Quantitative Research Topics For STEM Students interested in experimental quantitative research:
- Examining the impact of different fertilizers on crop yield in agriculture.
- Investigating the relationship between exercise and heart rate among different age groups.
- Analyzing the effect of varying light intensities on photosynthesis in plants.
- Studying the efficiency of various insulation materials in reducing building heat loss.
- Investigating the relationship between pH levels and the rate of corrosion in metals.
- Analyzing the impact of different concentrations of pollutants on aquatic ecosystems.
- Examining the effectiveness of different antibiotics on bacterial growth.
- Trying to figure out how temperature affects how thick liquids are.
- Finding out if there is a link between the amount of pollution in the air and lung illnesses in cities.
- Analyzing the efficiency of solar panels in converting sunlight into electricity under varying conditions.
Descriptive Research Topics for STEM Students
Descriptive research aims to provide a detailed account or description of a phenomenon. Here are 10 topics for STEM students interested in descriptive research:
- Describing the physical characteristics and behavior of a newly discovered species of marine life.
- Documenting the geological features and formations of a particular region.
- Creating a detailed inventory of plant species in a specific ecosystem.
- Describing the properties and behavior of a new synthetic polymer.
- Documenting the daily weather patterns and climate trends in a particular area.
- Providing a comprehensive analysis of the energy consumption patterns in a city.
- Describing the structural components and functions of a newly developed medical device.
- Documenting the characteristics and usage of traditional construction materials in a region.
- Providing a detailed account of the microbiome in a specific environmental niche.
- Describing the life cycle and behavior of a rare insect species.
Research Topics for STEM Students in the Pandemic:
The COVID-19 pandemic has raised many research opportunities for STEM students. Here are 10 research topics related to pandemics:
- Analyzing the effectiveness of various personal protective equipment (PPE) in preventing the spread of respiratory viruses.
- Studying the impact of lockdown measures on air quality and pollution levels in urban areas.
- Investigating the psychological effects of quarantine and social isolation on mental health.
- Analyzing the genomic variation of the SARS-CoV-2 virus and its implications for vaccine development.
- Studying the efficacy of different disinfection methods on various surfaces.
- Investigating the role of contact tracing apps in tracking & controlling the spread of infectious diseases.
- Analyzing the economic impact of the pandemic on different industries and sectors.
- Studying the effectiveness of remote learning in STEM education during lockdowns.
- Investigating the social disparities in healthcare access during a pandemic.
- Analyzing the ethical considerations surrounding vaccine distribution and prioritization.
Research Topics for STEM Students Middle School
Research topics for middle school STEM students should be engaging and suitable for their age group. Here are 10 research topics:
- Investigating the growth patterns of different types of mold on various food items.
- Studying the negative effects of music on plant growth and development.
- Analyzing the relationship between the shape of a paper airplane and its flight distance.
- Investigating the properties of different materials in making effective insulators for hot and cold beverages.
- Studying the effect of salt on the buoyancy of different objects in water.
- Analyzing the behavior of magnets when exposed to different temperatures.
- Investigating the factors that affect the rate of ice melting in different environments.
- Studying the impact of color on the absorption of heat by various surfaces.
- Analyzing the growth of crystals in different types of solutions.
- Investigating the effectiveness of different natural repellents against common pests like mosquitoes.
Technology Research Topics for STEM Students
Technology is at the forefront of STEM fields. Here are 10 research topics for STEM students interested in technology:
- Developing and optimizing algorithms for autonomous drone navigation in complex environments.
- Exploring the use of blockchain technology for enhancing the security and transparency of supply chains.
- Investigating the applications of virtual reality (VR) and augmented reality (AR) in medical training and surgery simulations.
- Studying the potential of 3D printing for creating personalized prosthetics and orthopedic implants.
- Analyzing the ethical and privacy implications of facial recognition technology in public spaces.
- Investigating the development of quantum computing algorithms for solving complex optimization problems.
- Explaining the use of machine learning and AI in predicting and mitigating the impact of natural disasters.
- Studying the advancement of brain-computer interfaces for assisting individuals with
- disabilities.
- Analyzing the role of wearable technology in monitoring and improving personal health and wellness.
- Investigating the use of robotics in disaster response and search and rescue operations.
Scientific Research Topics for STEM Students
Scientific research encompasses a wide range of topics. Here are 10 research topics for STEM students focusing on scientific exploration:
- Investigating the behavior of subatomic particles in high-energy particle accelerators.
- Studying the ecological impact of invasive species on native ecosystems.
- Analyzing the genetics of antibiotic resistance in bacteria and its implications for healthcare.
- Exploring the physics of gravitational waves and their detection through advanced interferometry.
- Investigating the neurobiology of memory formation and retention in the human brain.
- Studying the biodiversity and adaptation of extremophiles in harsh environments.
- Analyzing the chemistry of deep-sea hydrothermal vents and their potential for life beyond Earth.
- Exploring the properties of superconductors and their applications in technology.
- Investigating the mechanisms of stem cell differentiation for regenerative medicine.
- Studying the dynamics of climate change and its impact on global ecosystems.
Interesting Research Topics for STEM Students:
Engaging and intriguing research topics can foster a passion for STEM. Here are 10 interesting research topics for STEM students:
- Exploring the science behind the formation of auroras and their cultural significance.
- Investigating the mysteries of dark matter and dark energy in the universe.
- Studying the psychology of decision-making in high-pressure situations, such as sports or
- emergencies.
- Analyzing the impact of social media on interpersonal relationships and mental health.
- Exploring the potential for using genetic modification to create disease-resistant crops.
- Investigating the cognitive processes involved in solving complex puzzles and riddles.
- Studying the history and evolution of cryptography and encryption methods.
- Analyzing the physics of time travel and its theoretical possibilities.
- Exploring the role of Artificial Intelligence in creating art and music.
- Investigating the science of happiness and well-being, including factors contributing to life satisfaction.
Practical Research Topics for STEM Students
Practical research often leads to real-world solutions. Here are 10 practical research topics for STEM students:
- Developing an affordable and sustainable water purification system for rural communities.
- Designing a low-cost, energy-efficient home heating and cooling system.
- Investigating strategies for reducing food waste in the supply chain and households.
- Studying the effectiveness of eco-friendly pest control methods in agriculture.
- Analyzing the impact of renewable energy integration on the stability of power grids.
- Developing a smartphone app for early detection of common medical conditions.
- Investigating the feasibility of vertical farming for urban food production.
- Designing a system for recycling and upcycling electronic waste.
- Studying the environmental benefits of green roofs and their potential for urban heat island mitigation.
- Analyzing the efficiency of alternative transportation methods in reducing carbon emissions.
Experimental Research Topics for STEM Students About Plants
Plants offer a rich field for experimental research. Here are 10 experimental research topics about plants for STEM students:
- Investigating the effect of different light wavelengths on plant growth and photosynthesis.
- Studying the impact of various fertilizers and nutrient solutions on crop yield.
- Analyzing the response of plants to different types and concentrations of plant hormones.
- Investigating the role of mycorrhizal in enhancing nutrient uptake in plants.
- Studying the effects of drought stress and water scarcity on plant physiology and adaptation mechanisms.
- Analyzing the influence of soil pH on plant nutrient availability and growth.
- Investigating the chemical signaling and defense mechanisms of plants against herbivores.
- Studying the impact of environmental pollutants on plant health and genetic diversity.
- Analyzing the role of plant secondary metabolites in pharmaceutical and agricultural applications.
- Investigating the interactions between plants and beneficial microorganisms in the rhizosphere.
Qualitative Research Topics for STEM Students in the Philippines
Qualitative research in the Philippines can address local issues and cultural contexts. Here are 10 qualitative research topics for STEM students in the Philippines:
- Exploring indigenous knowledge and practices in sustainable agriculture in Filipino communities.
- Studying the perceptions and experiences of Filipino fishermen in coping with climate change impacts.
- Analyzing the cultural significance and traditional uses of medicinal plants in indigenous Filipino communities.
- Investigating the barriers and facilitators of STEM education access in remote Philippine islands.
- Exploring the role of traditional Filipino architecture in natural disaster resilience.
- Studying the impact of indigenous farming methods on soil conservation and fertility.
- Analyzing the cultural and environmental significance of mangroves in coastal Filipino regions.
- Investigating the knowledge and practices of Filipino healers in treating common ailments.
- Exploring the cultural heritage and conservation efforts of the Ifugao rice terraces.
- Studying the perceptions and practices of Filipino communities in preserving marine biodiversity.
Science Research Topics for STEM Students
Science offers a diverse range of research avenues. Here are 10 science research topics for STEM students:
- Investigating the potential of gene editing techniques like CRISPR-Cas9 in curing genetic diseases.
- Studying the ecological impacts of species reintroduction programs on local ecosystems.
- Analyzing the effects of microplastic pollution on aquatic food webs and ecosystems.
- Investigating the link between air pollution and respiratory health in urban populations.
- Studying the role of epigenetics in the inheritance of acquired traits in organisms.
- Analyzing the physiology and adaptations of extremophiles in extreme environments on Earth.
- Investigating the genetics of longevity and factors influencing human lifespan.
- Studying the behavioral ecology and communication strategies of social insects.
- Analyzing the effects of deforestation on global climate patterns and biodiversity loss.
- Investigating the potential of synthetic biology in creating bioengineered organisms for beneficial applications.
Correlational Research Topics for STEM Students
Correlational research focuses on relationships between variables. Here are 10 correlational research topics for STEM students:
- Analyzing the correlation between dietary habits and the incidence of chronic diseases.
- Studying the relationship between exercise frequency and mental health outcomes.
- Investigating the correlation between socioeconomic status and access to quality healthcare.
- Analyzing the link between social media usage and self-esteem in adolescents.
- Studying the correlation between academic performance and sleep duration among students.
- Investigating the relationship between environmental factors and the prevalence of allergies.
- Analyzing the correlation between technology use and attention span in children.
- Studying how environmental factors are related to the frequency of allergies.
- Investigating the link between parental involvement in education and student achievement.
- Analyzing the correlation between temperature fluctuations and wildlife migration patterns.
Quantitative Research Topics for STEM Students in the Philippines
Quantitative research in the Philippines can address specific regional issues. Here are 10 quantitative research topics for STEM students in the Philippines
- Analyzing the impact of typhoons on coastal erosion rates in the Philippines.
- Studying the quantitative effects of land use change on watershed hydrology in Filipino regions.
- Investigating the quantitative relationship between deforestation and habitat loss for endangered species.
- Analyzing the quantitative patterns of marine biodiversity in Philippine coral reef ecosystems.
- Studying the quantitative assessment of water quality in major Philippine rivers and lakes.
- Investigating the quantitative analysis of renewable energy potential in specific Philippine provinces.
- Analyzing the quantitative impacts of agricultural practices on soil health and fertility.
- Studying the quantitative effectiveness of mangrove restoration in coastal protection in the Philippines.
- Investigating the quantitative evaluation of indigenous agricultural practices for sustainability.
- Analyzing the quantitative patterns of air pollution and its health impacts in urban Filipino areas.
Things That Must Keep In Mind While Writing Quantitative Research Title
Here are a few things that must be kept in mind while writing a quantitative research:
1. Be Clear and Precise
Make sure your research title is clear and says exactly what your study is about. People should easily understand the topic and goals of your research by reading the title.
2. Use Important Words
Include words that are crucial to your research, like the main subjects, who you’re studying, and how you’re doing your research. This helps others find your work and understand what it’s about.
3. Avoid Confusing Words
Stay away from words that might confuse people. Your title should be easy to grasp, even if someone isn’t an expert in your field.
4. Show Your Research Approach
Tell readers what kind of research you did, like experiments or surveys. This gives them a hint about how you conducted your study.
5. Match Your Title with Your Research Questions
Make sure your title matches the questions you’re trying to answer in your research. It should give a sneak peek into what your study is all about and keep you on the right track as you work on it.
STEM students, addressing what STEM is and why research matters in this field. It offered an extensive list of research topics , including experimental, qualitative, and regional options, catering to various academic levels and interests. Whether you’re a middle school student or pursuing advanced studies, these topics offer a wealth of ideas. The key takeaway is to choose a topic that resonates with your passion and aligns with your goals, ensuring a successful journey in STEM research. Choose the best Experimental Quantitative Research Topics For Stem Students today!
Related Posts

Step by Step Guide on The Best Way to Finance Car

The Best Way on How to Get Fund For Business to Grow it Efficiently
Academia.edu no longer supports Internet Explorer.
To browse Academia.edu and the wider internet faster and more securely, please take a few seconds to upgrade your browser .
Enter the email address you signed up with and we'll email you a reset link.
- We're Hiring!
- Help Center

ADVANTAGES OF TAKING UP STEM IN SENIOR HIGH SCHOOL: PREPARATION FOR COLLEGE


Related Papers
Problems of education in the 21st century
Rita Birziņa
Philip Sadler
The Review of Higher Education
Catherine A Manly
2017 ASEE Annual Conference & Exposition Proceedings
Charles Lam
Karen Murcia
Science Education
Martha Bottia
Professional School Counseling
Proceedings of the 2014 ACM International Joint Conference on Pervasive and Ubiquitous Computing
learn everything
New Directions for Higher Education
Molly Schaller
Lfe Revista De Lenguas Para Fines Especificos
Loading Preview
Sorry, preview is currently unavailable. You can download the paper by clicking the button above.
RELATED PAPERS
Pastoral Care in Education
Stan Tucker
cristina rocha
Journal of Women and Minorities in Science and Engineering
Janelle Gagnon
B Lindsay Lowell
Proceedings of the ACM 2012 conference on Computer Supported Cooperative Work - CSCW '12
Duyên Nguyễn
Computers, Phones, and the InternetDomesticating Information Technology
Robert Kraut
International Journal of Advanced Research
ranjita uniyal
Revista de Pedagogie - Journal of Pedagogy
Olimpius Istrate
The International Journal of Learning: Annual Review
Sharn Donnison
2019 ASEE Annual Conference & Exposition Proceedings
Paris Svoronos
Education Sciences
khaleel ALarabi
Handbook of Research on Administration, Policy, and Leadership in Higher Education
Brooke Houck, Ph.D.
Sarah Lench
2006 Annual Conference & Exposition Proceedings
Oguz A . Soysal
Vestnik Tomskogo gosudarstvennogo universiteta
2012 ASEE Annual Conference & Exposition Proceedings
Clark Vangilder
2008 38th Annual Frontiers in Education Conference
James Collofello
MATEC Web of Conferences
LILIANA G POPESCU
Higher Education Pedagogies
Dawn Bennett
Edwin Varela
Journal of STEM Education: Innovations and Research
Anna Tan-Wilson
Journal of Baltic science education
Heba EL-Deghaidy
Vasudha Kilaru
Turkish Journal of Electrical Engineering Computer Sciences
Riadh Habash
- We're Hiring!
- Help Center
- Find new research papers in:
- Health Sciences
- Earth Sciences
- Cognitive Science
- Mathematics
- Computer Science
- Academia ©2024
- Write my thesis
- Thesis writers
- Buy thesis papers
- Bachelor thesis
- Master's thesis
- Thesis editing services
- Thesis proofreading services
- Buy a thesis online
- Write my dissertation
- Dissertation proposal help
- Pay for dissertation
- Custom dissertation
- Dissertation help online
- Buy dissertation online
- Cheap dissertation
- Dissertation editing services
- Write my research paper
- Buy research paper online
- Pay for research paper
- Research paper help
- Order research paper
- Custom research paper
- Cheap research paper
- Research papers for sale
- Thesis subjects
- How It Works
55 Brilliant Research Topics For STEM Students

Primarily, STEM is an acronym for Science, Technology, Engineering, and Mathematics. It’s a study program that weaves all four disciplines for cross-disciplinary knowledge to solve scientific problems. STEM touches across a broad array of subjects as STEM students are required to gain mastery of four disciplines.
As a project-based discipline, STEM has different stages of learning. The program operates like other disciplines, and as such, STEM students embrace knowledge depending on their level. Since it’s a discipline centered around innovation, students undertake projects regularly. As a STEM student, your project could either be to build or write on a subject. Your first plan of action is choosing a topic if it’s written. After selecting a topic, you’ll need to determine how long a thesis statement should be .
Given that topic is essential to writing any project, this article focuses on research topics for STEM students. So, if you’re writing a STEM research paper or write my research paper , below are some of the best research topics for STEM students.
List of Research Topics For STEM Students
Quantitative research topics for stem students, qualitative research topics for stem students, what are the best experimental research topics for stem students, non-experimental research topics for stem students, capstone research topics for stem students, correlational research topics for stem students, scientific research topics for stem students, simple research topics for stem students, top 10 research topics for stem students, experimental research topics for stem students about plants, research topics for grade 11 stem students, research topics for grade 12 stem students, quantitative research topics for stem high school students, survey research topics for stem students, interesting and informative research topics for senior high school stem students.
Several research topics can be formulated in this field. They cut across STEM science, engineering, technology, and math. Here is a list of good research topics for STEM students.
- The effectiveness of online learning over physical learning
- The rise of metabolic diseases and their relationship to increased consumption
- How immunotherapy can improve prognosis in Covid-19 progression
For your quantitative research in STEM, you’ll need to learn how to cite a thesis MLA for the topic you’re choosing. Below are some of the best quantitative research topics for STEM students.
- A study of the effect of digital technology on millennials
- A futuristic study of a world ruled by robotics
- A critical evaluation of the future demand in artificial intelligence
There are several practical research topics for STEM students. However, if you’re looking for qualitative research topics for STEM students, here are topics to explore.
- An exploration into how microbial factories result in the cause shortage in raw metals
- An experimental study on the possibility of older-aged men passing genetic abnormalities to children
- A critical evaluation of how genetics could be used to help humans live healthier and longer.
Experimental research in STEM is a scientific research methodology that uses two sets of variables. They are dependent and independent variables that are studied under experimental research. Experimental research topics in STEM look into areas of science that use data to derive results.
Below are easy experimental research topics for STEM students.
- A study of nuclear fusion and fission
- An evaluation of the major drawbacks of Biotechnology in the pharmaceutical industry
- A study of single-cell organisms and how they’re capable of becoming an intermediary host for diseases causing bacteria
Unlike experimental research, non-experimental research lacks the interference of an independent variable. Non-experimental research instead measures variables as they naturally occur. Below are some non-experimental quantitative research topics for STEM students.
- Impacts of alcohol addiction on the psychological life of humans
- The popularity of depression and schizophrenia amongst the pediatric population
- The impact of breastfeeding on the child’s health and development
STEM learning and knowledge grow in stages. The older students get, the more stringent requirements are for their STEM research topic. There are several capstone topics for research for STEM students .
Below are some simple quantitative research topics for stem students.
- How population impacts energy-saving strategies
- The application of an Excel table processor capabilities for cost calculation
- A study of the essence of science as a sphere of human activity
Correlations research is research where the researcher measures two continuous variables. This is done with little or no attempt to control extraneous variables but to assess the relationship. Here are some sample research topics for STEM students to look into bearing in mind how to cite a thesis APA style for your project.
- Can pancreatic gland transplantation cure diabetes?
- A study of improved living conditions and obesity
- An evaluation of the digital currency as a valid form of payment and its impact on banking and economy
There are several science research topics for STEM students. Below are some possible quantitative research topics for STEM students.
- A study of protease inhibitor and how it operates
- A study of how men’s exercise impacts DNA traits passed to children
- A study of the future of commercial space flight
If you’re looking for a simple research topic, below are easy research topics for STEM students.
- How can the problem of Space junk be solved?
- Can meteorites change our view of the universe?
- Can private space flight companies change the future of space exploration?
For your top 10 research topics for STEM students, here are interesting topics for STEM students to consider.
- A comparative study of social media addiction and adverse depression
- The human effect of the illegal use of formalin in milk and food preservation
- An evaluation of the human impact on the biosphere and its results
- A study of how fungus affects plant growth
- A comparative study of antiviral drugs and vaccine
- A study of the ways technology has improved medicine and life science
- The effectiveness of Vitamin D among older adults for disease prevention
- What is the possibility of life on other planets?
- Effects of Hubble Space Telescope on the universe
- A study of important trends in medicinal chemistry research
Below are possible research topics for STEM students about plants:
- How do magnetic fields impact plant growth?
- Do the different colors of light impact the rate of photosynthesis?
- How can fertilizer extend plant life during a drought?
Below are some examples of quantitative research topics for STEM students in grade 11.
- A study of how plants conduct electricity
- How does water salinity affect plant growth?
- A study of soil pH levels on plants
Here are some of the best qualitative research topics for STEM students in grade 12.
- An evaluation of artificial gravity and how it impacts seed germination
- An exploration of the steps taken to develop the Covid-19 vaccine
- Personalized medicine and the wave of the future
Here are topics to consider for your STEM-related research topics for high school students.
- A study of stem cell treatment
- How can molecular biological research of rare genetic disorders help understand cancer?
- How Covid-19 affects people with digestive problems
Below are some survey topics for qualitative research for stem students.
- How does Covid-19 impact immune-compromised people?
- Soil temperature and how it affects root growth
- Burned soil and how it affects seed germination
Here are some descriptive research topics for STEM students in senior high.
- The scientific information concept and its role in conducting scientific research
- The role of mathematical statistics in scientific research
- A study of the natural resources contained in oceans
Final Words About Research Topics For STEM Students
STEM topics cover areas in various scientific fields, mathematics, engineering, and technology. While it can be tasking, reducing the task starts with choosing a favorable topic. If you require external assistance in writing your STEM research, you can seek professional help from our experts.
Leave a Reply Cancel reply
Thank you for visiting nature.com. You are using a browser version with limited support for CSS. To obtain the best experience, we recommend you use a more up to date browser (or turn off compatibility mode in Internet Explorer). In the meantime, to ensure continued support, we are displaying the site without styles and JavaScript.
- View all journals
- Explore content
- About the journal
- Publish with us
- Sign up for alerts
- Open access
- Published: 30 August 2024
Unidirectional association of clonal hematopoiesis with atherosclerosis development
- Miriam Díez-Díez ORCID: orcid.org/0000-0003-1945-825X 1 na1 ,
- Beatriz L. Ramos-Neble ORCID: orcid.org/0000-0003-1479-8490 1 na1 ,
- Jorge de la Barrera 1 ,
- J. C. Silla-Castro 1 ,
- Ana Quintas ORCID: orcid.org/0000-0003-3721-7580 1 ,
- Enrique Vázquez 1 ,
- M. Ascensión Rey-Martín 1 ,
- Benedetta Izzi 1 ,
- Lucía Sánchez-García 1 ,
- Inés García-Lunar 1 , 2 , 3 ,
- Guiomar Mendieta 1 , 4 , 5 ,
- Virginia Mass 1 ,
- Nuria Gómez-López 1 ,
- Cristina Espadas 1 ,
- Gema González 1 ,
- Antonio J. Quesada ORCID: orcid.org/0000-0002-3280-8819 1 ,
- Ana García-Álvarez 1 , 2 , 4 , 5 , 6 ,
- Antonio Fernández-Ortiz 1 , 2 , 7 ,
- Enrique Lara-Pezzi 1 , 2 ,
- Ana Dopazo ORCID: orcid.org/0000-0002-4910-1684 1 , 2 ,
- Fátima Sánchez-Cabo ORCID: orcid.org/0000-0003-1881-1664 1 , 2 ,
- Borja Ibáñez ORCID: orcid.org/0000-0002-5036-254X 1 , 2 , 8 ,
- Vicente Andrés ORCID: orcid.org/0000-0002-0125-7209 1 , 2 ,
- Valentín Fuster ORCID: orcid.org/0009-0005-7344-8425 1 , 9 &
- José J. Fuster ORCID: orcid.org/0000-0002-5970-629X 1 , 2
Nature Medicine ( 2024 ) Cite this article
Metrics details
- Atherosclerosis
- Translational research
Clonal hematopoiesis, a condition in which acquired somatic mutations in hematopoietic stem cells lead to the outgrowth of a mutant hematopoietic clone, is associated with a higher risk of hematological cancer and a growing list of nonhematological disorders, most notably atherosclerosis and associated cardiovascular disease. However, whether accelerated atherosclerosis is a cause or a consequence of clonal hematopoiesis remains a matter of debate. Some studies support a direct contribution of certain clonal hematopoiesis-related mutations to atherosclerosis via exacerbation of inflammatory responses, whereas others suggest that clonal hematopoiesis is a symptom rather than a cause of atherosclerosis, as atherosclerosis or related traits may accelerate the expansion of mutant hematopoietic clones. Here we combine high-sensitivity DNA sequencing in blood and noninvasive vascular imaging to investigate the interplay between clonal hematopoiesis and atherosclerosis in a longitudinal cohort of healthy middle-aged individuals. We found that the presence of a clonal hematopoiesis-related mutation confers an increased risk of developing de novo femoral atherosclerosis over a 6-year period, whereas neither the presence nor the extent of atherosclerosis affects mutant cell expansion during this timeframe. These findings indicate that clonal hematopoiesis unidirectionally promotes atherosclerosis, which should help translate the growing understanding of this condition into strategies for the prevention of atherosclerotic cardiovascular disease in individuals exhibiting clonal hematopoiesis.
Somatic mutations that accumulate during the human lifespan are emerging as potential contributors to several disorders beyond their known role in cancer 1 . The most prominent example is that of clonal hematopoiesis (CH), a condition in which de novo somatic mutations in hematopoietic stem cells (HSCs) provide a selective advantage that leads to the clonal outgrowth of the mutant cell 2 , 3 . CH-associated mutations affect a limited set of genes—most frequently the epigenetic regulatory genes DNMT3A and TET2 —that are also commonly mutated in hematological neoplasia. Accordingly, CH is associated with a higher risk of developing hematological cancer 4 , 5 , 6 , 7 , 8 . In addition, CH is also emerging as a shared risk factor for a growing list of nonhematological disorders 4 , 6 , 7 , 9 , 10 , 11 , 12 , 13 , 14 , 15 , 16 , 17 , 18 , 19 , 20 , 21 , 22 , 23 . Atherosclerotic cardiovascular disease (CVD) is the nonhematological condition most strongly linked to CH 2 , 3 , 4 , 9 , 13 , 19 , 24 . However, whether accelerated atherosclerosis is a cause or a consequence of CH remains a matter of debate. While sequencing studies in humans and experimental studies in mice support a direct causal contribution of some CH-related mutations to atherosclerosis by exacerbating inflammatory responses 4 , 9 , 13 , 19 , 25 , 26 , mathematical models based on human and mouse data suggest that the association between CH and CVD may instead reflect reverse causality 27 . Specifically, it has been posited that CH could simply be a direct consequence of increased HSC proliferation in individuals with atherosclerosis, resulting in accelerated expansion of existing mutant hematopoietic clones 27 , 28 , 29 . This reverse causality hypothesis could also apply to other CH-related associations, thereby explaining why CH is associated with a very diverse range of conditions 4 , 6 , 7 , 9 , 10 , 11 , 12 , 13 , 14 , 15 , 16 , 17 , 18 , 30 , 31 , 32 . The uncertain nature of the relationship between CH and atherosclerosis has particularly important clinical implications, as the specific atherogenic effects of CH mutations on leukocytes are being discussed as potential targets for the development of personalized strategies to reduce or prevent CVD in individuals with CH 3 . Such approaches would not be effective if the association between CH and atherosclerosis is merely due to the effects of atherosclerosis on the dynamics of CH.
Resolving the controversy surrounding the directionality of the CH–atherosclerosis association requires longitudinal analyses that combine deep genotyping of CH-related genes with deep phenotyping of study participants. Such longitudinal analyses were not feasible in earlier studies, which mainly relied on cross-sectional whole-exome sequencing (WES) datasets at a single time point 4 , 6 , 7 , 9 , 13 , 16 . To address this issue, here we take advantage of the Progression of Early Subclinical Atherosclerosis (PESA) study 33 , a longitudinal cohort of healthy middle-aged individuals with serially collected blood DNA samples and deep cardiovascular and metabolic phenotyping, including extensive imaging-based assessment of subclinical atherosclerosis burden in multiple vascular territories and at multiple time points. Leveraging this resource, we used a high-sensitivity sequencing approach to investigate CH and its dynamics in a longitudinal manner, aiming to elucidate the interplay between CH and atherosclerosis.
CH in healthy middle-aged individuals
Previous analyses of WES or whole-genome sequencing (WGS) datasets suggested that CH is relatively uncommon in middle-aged individuals, with frequencies ranging approximately from 2% to 3% in individuals aged between 40 and 55 years, compared with >10% in individuals older than 65 (refs. 4 , 6 , 7 , 8 , 34 ). However, these previous observations were limited by the low sensitivity of somatic mutation calling based on WES or WGS data, which hampers the detection of small mutant clones (for example those present with variant allele fraction (VAF) <5%, that is, 10% of mutant blood cells, assuming monoallelic mutations) 34 . Therefore, to elucidate the prevalence and characteristics of CH in this segment of the population, we performed high-sensitivity targeted sequencing of 54 CH-related genes (median depth 3,712×; Extended Data Fig. 1a ) in blood samples collected at enrollment from 3,692 participants in PESA (40–55 years old at the time of enrollment, median 45; see Methods for details). We predefined a minimum VAF threshold of 0.2% to identify CH mutations, as our sequencing depth allowed us to detect CH variants at this VAF with a sensitivity greater than 90% ( Methods and Extended Data Fig. 1b ). Using this threshold and a previously described filtering and curation strategy 35 , 36 ( Methods and Supplementary Table 1 ), we identified a total of 1,172 CH-related somatic mutations in 46 genes, with 1,079 variants (92%) occurring in 33 genes linked to myeloid CH (typically referred to as CH of indeterminate potential or CHIP) and 93 variants in 13 genes linked to lymphoid CH 8 , 37 (Extended Data Fig. 1a and Supplementary Table 2 ). Approximately 25% of individuals carried at least one CH-related mutation detectable with our sequencing approach. Mutations in DNMT3A were most frequent (657 mutations, driving CH in 14.8% of individuals) followed by mutations in TET2 (153 mutations, driving CH in 3.9% of individuals), consistent with previous studies (Fig. 1a and Extended Data Fig. 1c ) 4 , 6 , 8 , 34 , 38 . The basic features of CH mutations in the study population aligned well with those reported in previous WES or WGS analyses 4 , 6 , 34 , 38 (Extended Data Fig. 1d and Supplementary Table 2 ), with more than one-half of the single nucleotide variants corresponding to a C > T substitution, a mutational signature characteristic of aging and CH (Extended Data Fig. 1e ).

We performed deep targeted sequencing to identify somatic mutations in a custom panel of 54 CH-related genes in 3,692 individuals from the PESA cohort. a , The number of CH driver mutations identified per gene. The values above the bars indicate the percentage of mutations affecting each specific gene. b , The CH prevalence across quartiles of age. c , The number of mutations per individual across quartiles of age. d , The association between advancing age (stratified as quartiles) and CH (analyzed separately as driven by mutations in DNMT3A , TET2 or other genes) based on multivariate logistic regression analyses adjusted for sex. The bars indicate 95% confidence intervals centered in the mean value (square). e , The distribution of mutant clone size in the study population, assessed as VAF. The dashed line shows the 2% VAF threshold most typically used to identify CH. The box shows the 25th (Q1), 50th (median) and 75th (Q3) percentiles of the data. The whiskers represent Q1 − 1.5 × IQR at the minimum and Q3 + 1.5 × IQR at the maximum. f , The prevalence of CH with VAF ≥2% across quartiles of age. g , The association between gene-specific CH and female sex, based on multivariate logistic regression analyses adjusted for age. The bars indicate 95% confidence intervals centered in the mean value (square). h , The CH prevalence across quartiles of age stratified by sex. In b , f and h , CH status in individuals carrying more than one mutation was defined on the basis of the mutation with the highest VAF.
The prevalence of CH mutations in this middle-aged population increased with advancing age (Fig. 1b ). After adjustment for sex, each additional year of age was independently associated with a 9% higher relative risk of carrying detectable CH mutations (odds ratio (OR) 1.09, 95% confidence interval (CI) 1.07–1.11, P < 0.001), and the prevalence of CH was 33.5% in the top quartile of age, which included individuals between ∼ 50 and ∼ 55 years of age (Fig. 1b ). Most mutation carriers (76%) had only one detectable mutation, although the frequency of carrying more than one mutation increased with age (Fig. 1c ). Age was markedly associated with DNMT3A -mutant CH (OR 1.09, 95% CI 1.07–1.12, P < 0.001), TET2 -mutant CH (OR 1.10, 95% CI 1.05–1.14, P < 0.001) and CH driven by mutations in other genes (OR 1.11, 95% CI 1.08–1.14, P < 0.001), although the increase in CH prevalence with aging differed between driver genes, with DNMT3A -mutant CH rising in prevalence earlier in life than CH driven by mutations in other genes (Fig. 1d ).
As expected, the mutant hematopoietic clones identified in our study were generally of much smaller size than those identified in previous WES or WGS analyses 4 , 6 , 34 , 38 (median VAF 0.92%, range 0.2–39.9%; Fig. 1e , Extended Data Fig. 1f and Supplementary Table 2 ). In total, 78.8% of the detected mutations had a VAF smaller than 2%, the minimum clone size threshold used in most previous CH studies 4 , 5 , 6 , 7 , 8 , 10 , 11 , 12 , 13 , 14 , 15 , 16 , 17 , and only ∼ 4% of them had a VAF greater than 10%, which identified clones with significant impact on atherosclerotic CVD in previous large studies 4 , 7 , 9 , 13 , 16 , 39 . CH mutations over the typical 2% VAF threshold were detected in 6.0% of the study population and in ∼ 9% of those between ∼ 50 and ∼ 55 years of age (Fig. 1f ). Computational downsampling of sequencing depth suggests that this elevated prevalence of CH compared with previous WGS/WES studies 4 , 6 , 7 , 8 , 34 reflects the better sensitivity provided by our targeted sequencing approach (Extended Data Fig. 1g,h ).
Consistent with recent reports 6 , 40 , we found that DNMT3A -mutant CH was associated with female sex after adjusting for age, with women exhibiting a 64% higher risk of carrying detectable DNMT3A mutations than men (Fig. 1g ). In total, 17.8% of women in the study population carried DNMT3A mutations, whereas 13.2% of men carried such mutations, and this higher frequency of DNMT3A mutations in women was evident across all quartiles of age (Fig. 1h ). No significant sex-related differences were observed in the prevalence of CH driven by mutations in genes other than DNMT3A (Fig. 1g,h ).
Clinical characteristics of CH mutation carriers
Having determined with high resolution the landscape of CH-related mutations in our study population, we next aimed to examine the pathophysiological effects of CH in these middle-aged individuals. Table 1 presents the demographic and clinical characteristics of the study population, stratified by CH status. The characteristics of individuals with clones with VAF ≥2% or gene-specific forms of CH are listed in Extended Data Table 1 and Supplementary Table 3 , respectively. Carrying CH-related mutations was associated with higher absolute blood cell counts in all hematopoietic lineages in multivariate regression analysis adjusted for age and sex (Extended Data Table 2 ). However, this effect was mild overall, with 3.9% and 7.2% relative increases in median counts of circulating leukocytes in those carrying CH-related mutations with VAF <2% and ≥2%, respectively (Extended Data Table 3 ). No other associations with major hematological parameters were observed. Overall, these findings corroborate the known relationship between CH and blood cell counts 6 , 7 , 13 , 34 . CH mutation carriers also tended to exhibit higher blood pressure and higher levels of glycated hemoglobin HbA1C (Table 1 ), consistent with some previous reports 6 , 13 , 41 , but these associations were not statistically significant after adjustment for age and sex (Supplementary Table 4 ).
No cross-sectional association between CH and atherosclerosis
We next investigated the cross-sectional relationship between CH and subclinical atherosclerosis burden assessed at enrollment through noninvasive vascular imaging across multiple vascular beds, including three-dimensional vascular ultrasound imaging (3DVUS) of carotid and femoral arteries, and computed tomography (CT)-based determination of coronary artery calcium scores (CACS). Although PESA participants who carried CH mutations exhibited greater subclinical atherosclerosis burden (Extended Data Fig. 2a–d ), no significant association was observed after adjusting for age and sex, or age, sex and conventional cardiovascular risk factors (Extended Data Table 4 ). Similar results were obtained in gene-specific analyses (Supplementary Table 5 ).
CH is associated with de novo femoral atherosclerosis
We next took advantage of the longitudinal nature of the PESA study and evaluated whether individuals who carried CH mutations at enrollment exhibited a higher risk of developing de novo atherosclerosis over subsequent years (Fig. 2a ). To do this, we leveraged the availability of 3DVUS imaging data for the carotid and femoral arteries, collected at enrollment and the 3-year and 6-year follow-up imaging examinations in PESA (Fig. 2b ). This imaging modality allows for the direct detection of atherosclerotic plaques, including the small plaques that are characteristic of early atherosclerosis and can be expected to develop in this timespan 42 , 43 . CH mutation carriers exhibited markedly higher rates of de novo femoral atherosclerosis development at the 3-year follow-up, which correlated with mutant clone size (Fig. 2c ). Similar results were observed when restricting this analysis to mutations in CHIP genes or specific frequently mutated genes (Fig. 2c ), or when evaluating de novo atherosclerosis development using imaging data from the 6-year follow-up (Fig. 2d ).

We investigated the association between CH at baseline and de novo development of femoral atherosclerosis ∼ 3 years and ∼ 6 years after enrollment among PESA participants who initially lacked detectable atherosclerosis in this region. 3DVUS imaging was used to determine the presence of femoral atherosclerosis. a , Summary of study design. b , Representative images from femoral atherosclerosis burden, assessed by 3DVUS, in an individual exhibiting de novo femoral atherosclerosis development (that is, absence of detectable atherosclerosis at the baseline evaluation (left) and plaque development at follow-up (right); scale bar, 5 mm). c , d , Incidence of de novo femoral atherosclerosis development at the 3-year follow-up ( c , n = 2,347) or the 6-year follow-up ( d , n = 2,214) in individuals free of CH (no CH) and in individuals exhibiting CH with VAF 0.2–2% or ≥2%. Incidence of de novo femoral atherosclerosis is also shown for myeloid CH (CHIP) or CH driven by specific CHIP genes. Statistical significance in the analyses of CH and CHIP was evaluated using univariate logistic regression models ( P for trends are shown). In gene-specific analyses, statistical significance was examined through proportion tests relative to the non-CHIP carriers group (* P < 0.05, ** P ≤ 0.01, *** P ≤ 0.001). e , f , The association between CH or CHIP and de novo femoral atherosclerosis development at the 3-year follow-up ( e , n = 2,347) and 6-year follow-up ( f , n = 2,214) based on multivariate logistic regression analyses. Statistical models were adjusted for age, sex, smoking, lipid-lowering treatment and the AUC of SBP, fasting glucose, LDL-C, BMI, CACS and global atherosclerotic plaque volume assessed by 3DVUS imaging; the bars indicate 95% confidence intervals centered in the mean value (square).
To account for the possible effects of the cumulative exposure to risk factors or the systemic burden of atherosclerosis 27 , 28 , 29 , we next evaluated the effects of CH in statistical models adjusted for the area under the curve (AUC) of conventional CV risk factors, CACS (assessed by CT) and global atherosclerotic plaque volume (assessed by 3DVUS), in addition to age and sex (see Methods for details). Using this approach, CH was independently associated with a 1.5-fold higher risk of developing de novo femoral atherosclerosis at the 3-year follow-up among participants who lacked detectable atherosclerosis in this vascular region at baseline (Fig. 2e ). The magnitude of this effect was dependent on mutant clone size, with smaller clones (VAF 0.2–2%) conferring a 1.4-fold higher risk of developing de novo femoral atherosclerosis, which did not reach statistical significance ( P = 0.072), and larger clones leading to a 2.1-fold higher risk, an effect comparable to that of traditional modifiable risk factors such as dyslipidemia or smoking (Extended Data Fig. 3a ). These associations were independent of blood cell counts (Extended Data Fig. 3b ). The effect of CH was primarily driven by mutations in genes related to myeloid CH or CHIP, which were associated with 1.6-fold greater risk of de novo femoral atherosclerosis (Fig. 2e and Extended Data Fig. 3c,d ). In gene-specific analyses, CH driven by mutations in DNMT3A , TET2 and other genes were all associated with higher rates of de novo femoral atherosclerosis (Fig. 2e ). The effect of CH mutations remained consistent after correction for multiple hypothesis testing (Supplementary Table 6 ). Similar, albeit milder, results were also obtained when evaluating femoral atherosclerosis development at the 6-year follow-up in PESA (Fig. 2f ). No association was observed between CH and atherosclerosis development in the carotid artery (Supplementary Fig. 1 ), a region where atherosclerosis seems to develop at later ages based on previous analyses in PESA and other cohorts with multiterritorial vascular imaging data 42 , 44 , 45 .
No effect of subclinical atherosclerosis on CH dynamics
Finally, we determined the longitudinal dynamics of CH in healthy middle-aged participants in PESA to examine its potential modulation by the presence or extent of atherosclerosis. To do this, we sequenced blood samples collected at baseline and ∼ 6 years after enrollment from 718 CH mutation carriers (Fig. 3a ). We then used the serially determined VAF measurements as a surrogate for clone size to calculate the annual relative expansion rates (AER) of CH mutations in blood (see details in Methods ). Among 602 CH-related mutations that were detected at both time points with comparable sequencing depth and quality (Supplementary Table 7 ), the median AER was 6.5% per year. CH-related mutations in DNMT3A (381 variants) and TET2 (80 variants) expanded at median AER of 7.0% per year and 7.6% per year, respectively (Fig. 3b ). Overall, mutations in epigenetic regulatory genes showed numerically slower expansion rates than genes encoding splicing regulators or involved in the DNA damage response (Supplementary Fig. 2 ), consistent with previous findings 46 , 47 , 48 . As a control, we also quantified the expansion of 271 nondeleterious somatic variants in CH-related genes identified at both time points in 223 individuals not carrying CH driver mutations. As expected, these neutral mutations generally did not show clear evidence of expansion within the timeframe of the study (median AER: 1.4% per year) (Fig. 3b ).

We monitored the expansion of 602 CH-related mutations through serial sequencing of blood samples from 494 individuals at baseline and ∼ 6 years after enrollment. Then, we investigated the association between subclinical atherosclerosis burden or related traits at baseline and the AER of the mutant hematopoietic clones. a , Summary of study design. b , The AER of mutations in any CH gene ( n = 602) or specifically in DNMT3A ( n = 381), TET2 ( n = 80) or other genes ( n = 141), compared with nondeleterious variants in CH-related genes ( n = 271). Statistical significance was determined by two-sided Mann–Whitney–Wilcoxon tests (** P ≤ 0.01, **** P ≤ 0.0001). c , The AER of CH-related mutations stratified by age quartiles ( n = 602). d , AER of CH-related mutations stratified by sex ( n = 602). e , The correlation between AER of CH-related mutations and baseline variant allele frequency (VAF). The P value for two-sided Pearson correlation is indicated. f – i , The AER of CH-related mutations in individuals with no detectable atherosclerosis and across tertiles of plaque burden, based on CACS ( f , n = 600), global plaque volume ( g , n = 578), carotid plaque volume ( h , n = 597) or femoral plaque volume ( i , n = 581) measured by 3DVUS. j , k , The AER of CH-related mutations stratified by conventional modifiable risk factors ( j , n = 602 for all variables, except for obesity, n = 601) and quartiles of hsCRP ( k , n = 600). Statistical significance in c , d and f – k was determined by mixed-effects models adjusted for baseline VAF ( d ) or sex and baseline VAF ( c and f – k ). The boxes in b – d and f – k represent the 25th (Q1), 50th (median) and 75th (Q3) percentiles of the data. The whiskers represent Q1 − 1.5 × IQR at the minimum and Q3 + 1.5 × IQR at the maximum.
The AER of CH-related mutations was comparable across quartiles of age, consistent with previous findings (Fig. 3c ) 48 . In contrast, a trend to faster expansion was observed in men (Fig. 3d ) and among variants with lower baseline VAF (Fig. 3e ). Importantly, the expansion of CH variants was not affected by the presence or the extent of subclinical atherosclerosis at baseline, assessed through various noninvasive imaging modalities (Fig. 3f–i and Table 2 ). Similarly, the cumulative burden of atherosclerosis over the timeframe of this longitudinal analysis, estimated as the AUC of several vascular imaging parameters, had no significant effect on the dynamics of CH (Table 2 ). In sensitivity analyses, we obtained comparable results when defining mutant cell expansion with two alternative mathematical approaches used in other studies of CH dynamics, namely the logarithm of the ratio of VAFs at both time points 49 and the exponential growth rate 50 (Table 2 ). Similarly, neither high-sensitivity C-reactive protein (hsCRP), a biomarker of systemic inflammation, nor several conventional cardiovascular risk factors, such as dyslipidemia, diabetes, smoking or obesity, were associated with significant changes in AER or other indicators of clonal expansion (Fig. 3j,k and Extended Data Table 5 ). Hypertension was associated with higher AER ( P = 0.048; Fig. 3j ), but this relationship was not statistically significant when evaluating other indicators of CH dynamics or the effect of baseline or cumulative systolic blood pressure (SBP) (Extended Data Table 5 ).
CH has emerged in recent years as a potential shared driver of blood cancer and several nonhematological conditions, most notably atherosclerotic CVD 2 , 3 . However, the accumulating reports of positive associations between CH and a diverse range of conditions and outcomes 4 , 6 , 7 , 9 , 10 , 11 , 12 , 13 , 14 , 15 , 16 , 17 , 18 , 19 , 20 , 21 , 22 , 23 , 30 , 32 also lend support to the possibility that CH is simply a marker of a shared pathophysiological feature of these conditions, rather than a causal factor. The association between CH and atherosclerosis, in particular, has become a matter of active debate 27 , 29 , hindering the translation of research findings into new strategies for the prevention of CVD in CH mutation carriers. In this context, our longitudinal assessment of the interplay between CH, atherosclerosis and related traits provides important new insights into the relationship between CH and atherosclerosis.
Most importantly, our findings shed light on the directionality of the association between CH and atherosclerosis. Previous research in both humans and mice has suggested that certain CH mutations directly contribute to the development of atherosclerotic CVD 4 , 9 , 13 , 25 , 26 . In contrast, recent mathematical models based on human and mouse data have proposed that the association between CH and CVD may instead reflect reverse causality, whereby CH would simply be a direct consequence of increased HSC proliferation in individuals with atherosclerosis, resulting in accelerated kinetics of mutant cell expansion 27 , 29 . This reverse causality hypothesis is also applicable to the association between CH and other conditions, such as diabetes 4 , 23 , 41 , known to increase HSC proliferation 51 . The direction of these associations remains uncertain, partly due to the fact that most of them were identified through post hoc analyses of cross-sectional sequencing datasets 4 , 7 , 9 , 10 , 11 , 12 , 13 , 14 , 15 , 16 , 17 , 18 , 30 , 31 , 32 , which provide only a static ‘snapshot’ of CH at a given time and, therefore, very limited insight into the dynamics of this condition and its association with disorders that develop subclinically over the course of years. In this context, our analysis of longitudinal sequencing and imaging data provides novel insights into the relationship between CH and atherosclerosis, demonstrating that: (1) carrying CH mutations at a given time confers a greater risk of developing femoral atherosclerosis over subsequent years, and (2) the presence or extent of subclinical atherosclerosis has no significant effect on the expansion of mutant hematopoietic clones. These findings argue strongly against the possibility that the association between CH and atherosclerosis is due to reverse causality, thus providing an essential prior step for the development of clinical trials that evaluate the efficacy of personalized preventive care strategies tailored to the effects of specific CH mutations. Our findings, however, do not rule out that certain cardiometabolic conditions may accelerate CH dynamics to some extent. Indeed, we found that hypertensive individuals exhibit a non-statistically significant trend to faster expansion of mutant cells, an observation that demands further exploration given the emerging evidence suggesting an impact of hypertension on the bone marrow niche and hematopoiesis 52 , 53 , 54 . In addition, we cannot rule out the possibility that clinically overt CVD (for example, acute ischemic heart disease) modifies the dynamics of CH. Although this possibility would not affect the primary conclusions of our study, it deserves further investigation as it may help understand the regulation and effects of CH in high-risk CVD patients. Overall, further studies are warranted to dissect the factors that determine the dynamics of CH and explore the underlying regulatory mechanisms.
Our study also provides a large high-sensitivity assessment of the prevalence of CH and the magnitude of its connection to atherosclerosis. Previous human genetic evidence linking CH to atherosclerotic CVD was mainly obtained by repurposing WES datasets 4 , 7 , 9 , 13 , 16 , 39 , which were not intended initially to investigate somatic variants and have limited sensitivity to detect CH due to their modest sequencing depth. A previous report suggested that the typical depth of WES analyses only provides robust ability to identify mutant clones with VAF >5%, (that is, carriers of >10% mutant blood cells, if carrying monoallelic mutations) 34 , which is consistent with our estimations based on downsampling of sequencing depth in our study population. Furthermore, slight differences in variant interpretation and filtering parameters can be a source of major variability when using WES data to detect CH, as illustrated by recent studies by independent groups, who reached substantially different conclusions about the prevalence of CH and the strength of its association with atherosclerotic CVD despite analyzing WES data from the same cohort 7 , 8 , 16 . Here, through high-sensitivity sequencing of almost 3,700 seemingly healthy individuals, we demonstrate that CH mutations are prevalent in middle-aged individuals, being detectable in ∼ 25% of healthy individuals between 40 and 55 years of age, and in ∼ 34% of those aged 50–55. Even when applying the commonly used 2% VAF threshold for CH detection, we found that 6% of the study population (age range 40–55 years old) and 9% of those aged 50–55 exhibit CH, indicating an approximately three times higher prevalence of CH in middle-aged individuals than previously reported 4 , 6 , 7 , 8 , 34 . Thus, our findings suggest that CH contributes to atherosclerosis development in a much greater proportion of the population than previously believed. Importantly, the vast majority of mutant clones in our study population were small, with 79% of the mutations exhibiting a VAF <2% and 96% of them exhibiting a VAF <10% at baseline. Yet, carrying clones in this size range was still pathophysiologically relevant, as it was associated with a higher risk of de novo femoral atherosclerosis development, particularly among individuals with clones expanded to VAF ≥2%. These findings contribute to clarify the threshold of VAF that defines pathophysiologically relevant CH, as large WES-based studies previously concluded that clones with VAF <10% had minimal impact on subclinical atherosclerosis, atherosclerotic CVD or CVD-related mortality 6 , 7 , 9 , 16 , 39 , in contrast to smaller studies using more sensitive targeted sequencing approaches 19 .
Our results also provide important insight into gene-specific effects of CH. While the effect of somatic TET2 mutations on atherosclerotic CVD is supported by several sequencing studies 9 , 13 , 16 , the effect of DNMT3A mutations has remained controversial, with previous WES-based studies reporting a minimal, almost null effect of these mutations on atherosclerotic CVD 7 , 13 , 16 . In this context, while additional studies are warranted, our gene-specific analysis reinforces the association between somatic DNMT3A mutations and atherosclerotic disease, consistent with experiments in mouse models 55 and recent targeted sequencing studies 56 , and in contrast to WES-based analyses 7 , 13 , 16 . As DNMT3A mutations are the most frequent mutations linked to CH, these findings underscore the potential clinical significance of this condition.
Several limitations must be considered when interpreting our findings. First, the study population consisted entirely of Caucasian individuals residing in Madrid, Spain, which may limit the generalizability of our results to other racial and ethnic groups. Second, our findings related to the effects of CH on de novo femoral atherosclerosis cannot be directly extrapolated to other vascular beds; in this context, future studies with other imaging modalities and in other age ranges are warranted. Third, CH in this middle-aged population was markedly dominated by DNMT3A mutations, which limits our statistical power to assess the effects of CH driven by mutations in other genes; thus, gene-specific analyses in this work should be considered exploratory. Finally, our study was focused on examining the effects of atherosclerosis on the expansion of existing mutant hematopoietic clones; therefore, we cannot draw any conclusions regarding the potential effects of atherosclerosis or related traits on de novo somatic mutagenesis. Testing this possibility, which to our knowledge has not been postulated so far, would require more sensitive sequencing techniques, such as duplex sequencing.
In summary, our study elucidates the directionality of the association between CH and atherosclerosis in humans. These results provide critical new insight into the effects and regulation of CH, which should help translate our growing knowledge of this condition into personalized strategies for managing the risk of atherosclerotic CVD and, potentially, that of other age-related disorders.
Study population
The study population included 3,692 participants in the PESA study ( NCT01410318 ) with available blood DNA samples, multimodal vascular imaging data at multiple time points and written informed consent for DNA sequencing analyses. PESA is an ongoing observational prospective cohort study that is examining several imaging, biological and behavioral parameters potentially related to the presence and progression of early subclinical atherosclerosis in 4,184 apparently healthy middle-aged individuals. PESA participants were between 40 and 55 years old at enrollment (between June 2010 and February 2014), and have undergone three study visits so far (baseline and ∼ 3- and ∼ 6-year follow-up). All study visits included a clinical interview, physical examination, collection of blood samples and assessment of subclinical atherosclerosis by noninvasive imaging techniques, among other measurements 33 . Exclusion criteria included known CVD, cancer or immunological disorders, morbid obesity, chronic kidney disease, the presence of any disease expected to decrease life expectancy, and any condition that could affect adherence to the study procedures. The study protocol was approved by the Ethics Committee of Instituto de Salud Carlos III (CEI PI_52/2019), and all participants provided written informed consent.
Assessment of burden and progression of subclinical atherosclerosis and related traits
Atherosclerosis burden in the study population was assessed by noninvasive vascular imaging tests, in accordance with standard protocols in PESA, including 3DVUS-based assessment of the femoral and carotid arteries and noncontrast CT to determine CACS 33 , 42 , 43 , 44 . Individual-level 3DVUS data were collected using VPQ software v13 (Philips Healthcare) and CM2020 software (Philips Research Analysis). Atherosclerosis by 3DVUS was analyzed separately in carotid and femoral arteries as well as in an integrated manner (that is, global atherosclerotic plaque volume). Presence of subclinical atherosclerosis at baseline was defined as CACS >0 or plaque volume by 3DVUS >0 mm 3 (categorical variables). The extent of subclinical atherosclerosis at baseline was estimated on the basis of CACS and plaque volumes, either as continuous variables or categorized as 0 and tertiles. Representative 3DVUS images across tertiles of atherosclerotic plaque volume can be found in Extended Data Fig. 4 . In longitudinal analyses, de novo atherosclerosis development was defined as transitioning from absence of detectable atherosclerosis by 3DVUS at baseline to detectable atherosclerosis with plaque volume >0 mm 3 at follow-up. Limited sensitivity and statistical power precluded an accurate examination of the effects of CH on the progression of prevalent atherosclerotic plaques. Modifiable risk factors and related traits were defined as continuous or categorical variables and evaluated on the basis of analyses of fasting blood samples and questionnaires. Diabetes was defined as exhibiting plasma fasting glucose ≥126 mg dl −1 or being treated with insulin or hypoglycemic medication. Hypertension was defined as exhibiting SBP ≥140 mmHg or diastolic blood pressure (DBP) ≥90 mmHg, or being treated with antihypertensive medication. Dyslipidemia was defined as exhibiting total cholesterol ≥240 mg dl −1 , low-density lipoprotein cholesterol (LDL-C) ≥160 mg dl −1 or high-density lipoprotein cholesterol (HDL-C) <40 mg dl −1 , or being treated with lipid-lowering drugs. Obesity was defined as exhibiting a body mass index (BMI) ≥30 kg m −2 . Low plasma HDL-C was defined as exhibiting plasma HDL-C <40 mg dl −1 in males or <50 mg dl −1 in females. Hematological analyses included counts of leukocytes and main leukocyte subsets (neutrophils, lymphocytes, monocytes, eosinophils and basophils), red blood cell count, hemoglobin, hematocrit, red blood cell distribution width and platelet counts.
High-sensitivity targeted sequencing to identify CH-related somatic mutations
Sample preparation and high-sensitivity DNA sequencing, as well as somatic variant calling and annotation, were performed as previously described 35 , 36 . In brief, a custom gene panel was designed to identify somatic mutations in 54 well-established CH-related genes (Extended Data Fig. 1 and Supplementary Table 1 ), and unique dual indexes and unique molecular identifiers were used to detect polymerase chain reaction and sequencing errors and, therefore, to improve identification of small mutant clones. After confirming the quality of blood DNA, DNA libraries were generated (KAPA Hyper plus kit, Roche) and genes of interest were captured using xGen Hybridization Capture reagents (Integrated DNA Technologies). Then, DNA libraries were paired-end sequenced on a HiSeq 4000 or a NovaSeq 6000 sequencing system (Illumina). Raw sequencing reads were mapped to human genome build GRCh38, and somatic variants were identified by using Genome Analysis Toolkit (GATK) Mutect2. We removed common sequencing artifacts according to several Mutect2 filters (multiallelic somatic calls and variants with deficient mapping or base quality) as well as those flagged by the position filter. Germline mutations were excluded, and we only considered variants with ‘high’ or ‘moderate’ impact according to Variant Effect Predictor, reference read depth ≥300, alternative depth ≥4 and evidence of the variant on both forward and reverse strands (F1R2 and F2R1 ≥2). We also filtered out variants preceded or followed by homopolymer regions, indels with ≥3 consecutive repeated nucleotides within the insertion or deletion, and those SNVs located in a ≥5-nucleotide-long homopolymer, except if found at multiple sequencing time points in the same individual. Candidate CH driver mutations were then identified on the basis of prespecified inclusion criteria (Supplementary Table 1 ), the presence of ≥3 times in the Catalogue of Somatic Mutations in Cancer (COSMIC, https://cancer.sanger.ac.uk/cosmic ) in hematopoietic samples from at least three different studies, in silico pathogenicity predictors and previous publications using a similar deep sequencing strategy. A specific approach was required to identify mutations in U2AF1 57 because of an erroneous duplication in the region of this gene in the GRCh38reference genome 58 .
We set a minimum VAF threshold of 0.2% to identify CH mutations, as our sequencing depth allowed us to detect CH variants at this VAF with a sensitivity greater than 90% (Extended Data Fig. 1b ). In brief, we used a binomial distribution to estimate the sensitivity for the detection of somatic variants with a certain VAF at different sequencing depths, requiring a minimum alternative allele count of four reads. CH-related variants over the 0.2% VAF threshold were significantly associated with age (Supplementary Table 8 ), an essential feature of true CH variants 8 .
Assessment of expansion rates of mutant hematopoietic clones
Blood samples collected ∼ 6 years after enrollment from 718 CH mutation carriers were sequenced as described above. Among the 931 CH-related variants with VAF ≥0.2% identified in these individuals at baseline, 261 variants were not detected at follow-up. These variants were excluded from further analysis, as we were unable to ascertain if the apparent disappearance of these mutant clones is due to biological or technical reasons. We also excluded 68 additional variants for which sequencing depths differed substantially between both time points (ratio of baseline depth to follow-up depth exceeding the 90% percentile of the distribution). After applying this filtering strategy, 602 CH-related mutations in 494 individuals were detected at both time points supported by reads in both the forward and reverse strand (Supplementary Table 7 ). The AER of each of these mutations was calculated as the relative change in VAF over the follow-up period in years \((\Delta {{\mathrm{VAF}}} / {{\mathrm{VA}}}{{{{\mathrm{F}}}}}_{{\rm{baseline}}})/{{\mathrm{time}}}\times 100\) . As a sensitivity analysis, we also calculated two additional indicators of the expansion rate of mutant hematopoietic clones based on previous literature: the logarithmic ratio of VAFs, that is log(VAF/VAF baseline )/time × 100 (ref. 49 ), and the exponential growth rate, that is, VAF = VAF baseline × (clonal growth + 1) time ; clonal growth = ((VAF/VAF baseline ) (1/(time) ) − 1) × 100 (ref. 50 ). Time refers to the follow-up period measured in years. These three metrics showed a perfect positive correlation with each other, with Spearman’s rank correlation coefficients equal to 1.00 (Supplementary Fig. 3 ).
As a control, blood samples from 223 individuals not carrying CH-related mutations at baseline were also sequenced ∼ 6 years after enrollment to estimate the expansion of nondeleterious somatic variants in CH-related genes, as a surrogate of the expansion related to stochastic neutral drift. We considered nondeleterious somatic variants those with impact ‘low’ (unlikely to change protein behavior, for example, synonymous variants) or ‘modifier‘ (typically noncoding variants). Likely germline variants and common sequencing artifacts were filtered out, and only variants supported by reads on both the forward and reverse strands at both time points were included in the analysis, consistent with our approach to investigate the expansion of CH driver variants.
Statistical analysis
All statistical analyses were performed using RStudio (v2022.07.2 + 576) with R version 4.2.2, considering a statistical significance level of 0.05. Adjustments for multiple hypothesis testing were not performed unless otherwise indicated. Variables were expressed as frequency (%), mean and standard deviation, or median and interquartile range (IQR), as appropriate. CH was investigated as a composite of mutations in any CH-related gene, unless otherwise stated. Specific analyses evaluated the effects of mutations in DNMT3A , TET2 and other genes separately. Differences in baseline characteristics between CH mutation carriers and individuals with no detectable CH mutation were evaluated through independent two-tailed t -tests for continuous variables exhibiting a normal distribution, Wilcoxon rank tests for continuous variables with a nonnormal distribution and χ 2 tests for categorical variables. Associations between CH and traits of interest were evaluated using univariate or multivariate linear or logistic regression models, adjusted for age and sex or age, sex, LDL-C, SBP, BMI, fasting glucose levels, lipid-lowering treatment and smoking, as specified in figure legends. Some specific analyses were adjusted for age, sex and cardiovascular risk factors defined as categorical variables, as indicated in figure legends. Analyses of the association between CH and de novo development of atherosclerosis were adjusted by the AUCs of LDL-C, SBP, BMI and fasting glucose to consider the cumulative exposure to this cardiovascular risk factors during follow-up. These analyses were also adjusted for the AUC of CACS and global plaque volume to account for the possibility that the possible cumulative effects of systemic atherosclerosis on CH 27 confound its relationship with de novo atherosclerosis development in specific regions. In multivariate analyses, only individuals with information for all the covariates included in the model were considered. In analyses concerning specific types of CH (for example, driven by mutations in a given gene or over a certain VAF), individuals not carrying detectable CH mutations were used as the reference category. In individuals carrying more than one CH mutation, all mutations were considered separately in gene-specific analyses, unless otherwise stated. In longitudinal analyses of CH dynamics, differences in the AER of gene-specific mutations compared with the expansion of nondeleterious variants were evaluated using two-tailed Mann–Whitney–Wilcoxon tests. The association between variables of interest and mutant clone expansion was evaluated through mixed-effects models (R package lme4 v1.1.34). Based on an estimation of statistical power (using the G*power software v3.1.9.7 and the R package pwrss v0.3.1), we had >80% power to detect as statistically significant ( α = 0.05) any effect of the presence of subclinical atherosclerosis leading to an absolute increase in AER greater than 3.2%, an effect size substantially smaller than that expected to result from the increased HSC proliferation reported in human individuals with atherosclerosis 27 . To prevent potential biases, the analysis of imaging data, the sequencing of DNA samples, the bioinformatic processing of sequencing data, the curation of somatic variants to identify CH driver mutations and the statistical analysis of associations were conducted independently by different investigators who were blinded to other data.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
CH-related mutations identified through high-sensitivity genomic DNA sequencing are listed in Supplementary Table 2 . Other data from PESA participants are not publicly available for privacy and legal reasons. Access to these data can be requested to the PESA Scientific Committee via the corresponding authors or by e-mailing [email protected]. Data access will require a research proposal and approval by the PESA Scientific Committee, which meets every other month to evaluate such requests. To gain access, data requestors will need to sign a data access agreement and a nondisclosure agreement. Human genome build GRCh38 ( GCF_000001405.40 ) is available at https://gatk.broadinstitute.org/hc/en-us/articles/360035890811-Resource-bundle . The present article includes all other data generated or analyzed during this study.
Code availability
The code is publicly available and can be found via GitHub at https://github.com/Unidad-Bioinformatica-CNIC/CHIP-candidate_mutations . The source code from the R-packages used in this study are freely available online ( https://cran.r-project.org/ ).
Mustjoki, S. & Young, N. S. Somatic mutations in “benign” disease. N. Engl. J. Med. 384 , 2039–2052 (2021).
Article CAS PubMed Google Scholar
Jaiswal, S. & Ebert, B. L. Clonal hematopoiesis in human aging and disease. Science 366 , 6465 (2019).
Article Google Scholar
Tall, A. R. & Fuster, J. J. Clonal hematopoiesis in cardiovascular disease and therapeutic implications. Nat. Cardiovasc. Res. 1 , 116–124 (2022).
Article PubMed PubMed Central Google Scholar
Jaiswal, S. et al. Age-related clonal hematopoiesis associated with adverse outcomes. N. Engl. J. Med. 371 , 2488–2498 (2014).
Genovese, G. et al. Clonal hematopoiesis and blood-cancer risk inferred from blood DNA sequence. N. Engl. J. Med. 371 , 2477–2487 (2014).
Kar, S. P. et al. Genome-wide analyses of 200,453 individuals yield new insights into the causes and consequences of clonal hematopoiesis. Nat. Genet. 54 , 1155–1166 (2022).
Article CAS PubMed PubMed Central Google Scholar
Kessler, M. D. et al. Common and rare variant associations with clonal haematopoiesis phenotypes. Nature 612 , 301–309 (2022).
Vlasschaert, C. et al. A practical approach to curate clonal hematopoiesis of indeterminate potential in human genetic datasets. Blood 141 , 2214–2223 (2023).
CAS PubMed PubMed Central Google Scholar
Jaiswal, S. et al. Clonal hematopoiesis and risk of atherosclerotic cardiovascular disease. N. Engl. J. Med. 377 , 111–121 (2017).
Miller, P. G. et al. Association of clonal hematopoiesis with chronic obstructive pulmonary disease. Blood 139 , 357–368 (2022).
Agrawal, M. et al. TET2-mutant clonal hematopoiesis and risk of gout. Blood 140 , 1094–1103 (2022).
Kim, P. G. et al. Dnmt3a-mutated clonal hematopoiesis promotes osteoporosis. J. Exp. Med. 218 , e20211872 (2021).
Zekavat, S. M. et al. TP53-mediated clonal hematopoiesis confers increased risk for incident atherosclerotic disease. Nat. Cardiovasc. Res. 2 , 144–158 (2023).
Shi, C. et al. Clonal haematopoiesis of indeterminate potential: associations with heart failure incidence, clinical parameters and biomarkers. Eur. J. Heart Fail. 25 , 4–13 (2023).
Yu, B. et al. Supplemental association of clonal hematopoiesis with incident heart failure. J. Am. Coll. Cardiol. 78 , 42–52 (2021).
Vlasschaert, C., Heimlich, J. B., Rauh, M. J., Natarajan, P. & Bick, A. G. Interleukin-6 receptor polymorphism attenuates clonal hematopoiesis-mediated coronary artery disease risk among 451 180 individuals in the UK Biobank. Circulation 147 , 358–360 (2023).
Kestenbaum, B. et al. Clonal hematopoiesis of indeterminate potential and kidney function decline in the general population. Am. J. Kidney Dis. 81 , 329–335 (2023).
Wong, W. J. et al. Clonal haematopoiesis and risk of chronic liver disease. Nature 616 , 747–754 (2023).
Zhao, K. et al. Somatic and germline variants and coronary heart disease in a Chinese population. JAMA Cardiol. 9 , 233–242 (2024).
Article PubMed Google Scholar
Vlasschaert, C. et al. Clonal hematopoiesis of indeterminate potential is associated with acute kidney injury. Nat. Med. 30 , 810–817 (2024).
Schuermans, A. et al. Clonal haematopoiesis of indeterminate potential predicts incident cardiac arrhythmias. Eur. Heart J. 45 , 791–805 (2024).
Ahn, H. J. et al. Clonal haematopoiesis of indeterminate potential and atrial fibrillation: an East Asian cohort study. Eur. Heart J. 45 , 778–790 (2024).
Tobias, D. K. et al. Clonal hematopoiesis of indeterminate potential (CHIP) and incident type 2 diabetes risk. Diabetes Care 46 , 1978–1985 (2023).
Libby, P. et al. Clonal hematopoiesis: crossroads of aging, cardiovascular disease, and cancer: JACC review topic of the week. J. Am. Coll. Cardiol. 74 , 567–577 (2019).
Fuster, J. J. et al. Clonal hematopoiesis associated with TET2 deficiency accelerates atherosclerosis development in mice. Science 355 , 842–847 (2017).
Fidler, T. P. et al. The AIM2 inflammasome exacerbates atherosclerosis in clonal haematopoiesis. Nature 592 , 296–301 (2021).
Heyde, A. et al. Increased stem cell proliferation in atherosclerosis accelerates clonal hematopoiesis. Cell 184 , 1348–1361 e1322 (2021).
McAlpine, C. S. et al. Sleep exerts lasting effects on hematopoietic stem cell function and diversity. J. Exp. Med. 219 , e20220081 (2022).
Sanchez-Cabo, F. & Fuster, J. J. Clonal haematopoiesis and atherosclerosis: a chicken or egg question? Nat. Rev. Cardiol. 18 , 463–464 (2021).
Honigberg, M. C. et al. Premature menopause, clonal hematopoiesis, and coronary artery disease in postmenopausal women. Circulation 143 , 410–423 (2021).
Bohme, M. et al. Impact of clonal hematopoiesis in patients with cardiogenic shock complicating acute myocardial infarction. J. Am. Coll. Cardiol. 80 , 1545–1556 (2022).
Dharan, N. J. et al. HIV is associated with an increased risk of age-related clonal hematopoiesis among older adults. Nat. Med. 27 , 1006–1011 (2021).
Ibanez, B. et al. Progression of Early Subclinical Atherosclerosis (PESA) study: JACC Focus Seminar 7/8. J. Am. Coll. Cardiol. 78 , 156–179 (2021).
Bick, A. G. et al. Inherited causes of clonal haematopoiesis in 97,691 whole genomes. Nature 586 , 763–768 (2020).
Diez-Diez, M. et al. Clonal hematopoiesis is not prevalent in Hutchinson–Gilford progeria syndrome. Geroscience 45 , 1231–1236 (2023).
Pascual-Figal, D. A. et al. Clonal hematopoiesis and risk of progression of heart failure with reduced left ventricular ejection fraction. J. Am. Coll. Cardiol. 77 , 1747–1759 (2021).
Niroula, A. et al. Distinction of lymphoid and myeloid clonal hematopoiesis. Nat. Med. 27 , 1921–1927 (2021).
Buscarlet, M. et al. DNMT3A and TET2 dominate clonal hematopoiesis and demonstrate benign phenotypes and different genetic predispositions. Blood 130 , 753–762 (2017).
Bick, A. G. et al. Genetic interleukin 6 signaling deficiency attenuates cardiovascular risk in clonal hematopoiesis. Circulation 141 , 124–131 (2020).
Kamphuis, P. et al. Sex differences in the spectrum of clonal hematopoiesis. Hemasphere 7 , e832 (2023).
Kim, M. J. et al. Clonal hematopoiesis as a novel risk factor for type 2 diabetes mellitus in patients with hypercholesterolemia. Front. Public Health 11 , 1181879 (2023).
Lopez-Melgar, B. et al. Subclinical atherosclerosis burden by 3D ultrasound in mid-life: the PESA study. J. Am. Coll. Cardiol. 70 , 301–313 (2017).
Lopez-Melgar, B. et al. Accurate quantification of atherosclerotic plaque volume by 3D vascular ultrasound using the volumetric linear array method. Atherosclerosis 248 , 230–237 (2016).
Fernandez-Friera, L. et al. Prevalence, vascular distribution, and multiterritorial extent of subclinical atherosclerosis in a middle-aged cohort: the PESA (Progression of Early Subclinical Atherosclerosis) study. Circulation 131 , 2104–2113 (2015).
Laclaustra, M. et al. Femoral and carotid subclinical atherosclerosis association with risk factors and coronary calcium: the AWHS study. J. Am. Coll. Cardiol. 67 , 1263–1274 (2016).
Robertson, N. A. et al. Longitudinal dynamics of clonal hematopoiesis identifies gene-specific fitness effects. Nat. Med. 28 , 1439–1446 (2022).
Fabre, M. A. et al. The longitudinal dynamics and natural history of clonal haematopoiesis. Nature 606 , 335–342 (2022).
van Zeventer, I. A. et al. Evolutionary landscape of clonal hematopoiesis in 3,359 individuals from the general population. Cancer Cell 41 , 1017–1031 e1014 (2023).
Bolton, K. L. et al. Cancer therapy shapes the fitness landscape of clonal hematopoiesis. Nat. Genet. 52 , 1219–1226 (2020).
Mack, T. et al. Cost-effective and scalable clonal hematopoiesis assay provides insight into clonal dynamics. J. Mol. Diagn. 26 , 563–573 (2024).
Hoyer, F. F. et al. Bone marrow endothelial cells regulate myelopoiesis in diabetes mellitus. Circulation 142 , 244–258 (2020).
Devesa, A. et al. Bone marrow activation in response to metabolic syndrome and early atherosclerosis. Eur. Heart J. 43 , 1809–1828 (2022).
Al-Sharea, A. et al. Chronic sympathetic driven hypertension promotes atherosclerosis by enhancing hematopoiesis. Haematologica 104 , 456–467 (2019).
Rohde, D. et al. Bone marrow endothelial dysfunction promotes myeloid cell expansion in cardiovascular disease. Nat. Cardiovasc. Res. 1 , 28–44 (2022).
Rauch, P. J. et al. Loss-of-function mutations in Dnmt3a and Tet2 lead to accelerated atherosclerosis and concordant macrophage phenotypes. Nat. Cardiovasc. Res. 2 , 805–818 (2023).
Google Scholar
Wang, S. et al. Prevalence and prognostic significance of DNMT3A- and TET2- clonal haematopoiesis-driver mutations in patients presenting with ST-segment elevation myocardial infarction. EBioMedicine 78 , 103964 (2022).
Weinstock, J. S. et al. Aberrant activation of TCL1A promotes stem cell expansion in clonal haematopoiesis. Nature 616 , 755–763 (2023).
Miller, C. A. et al. Failure to detect mutations in U2AF1 due to changes in the GRCh38 reference sequence. J. Mol. Diagn. 24 , 219–223 (2022).
Download references
Acknowledgements
The PESA study is funded by the Centro Nacional de Investigaciones Cardiovasculares Carlos III (CNIC), Madrid, Spain, and Banco Santander, Madrid, Spain. The project leading to these results received funding from ‘la Caixa’ Foundation under the project codes LCF/PR/HR17/52150007 and LCF/PR/HR22/52420011. This research work was also supported by grant PLEC2021-008194, funded by MICIU/AEI/10.13039/501100011033 and by ‘European Union NextGenerationEU/PRTR’; grant PID2021-126580OB-I00, funded by MICIU/AEI/10.13039/501100011033 and by ERDF/EU; and grant 202314-31, funded by Fundació ‘La Marató TV3’. M.D.D. is supported by grant PRE2019-087463, funded by MICIU/AEI/10.13039/501100011033’ and ‘ESF Investing in your future’. B.L.R.-N. is supported by grant PRE2019-087829, funded by MICIU/AEI/10.13039/501100011033’ and ‘ESF Investing in your future’. B.Ib. is supported by the European Commission (grant nos. ERC-CoG 819775 and H2020-HEALTH 945118), the Spanish Ministry of Science and Innovation (PID2022-140176OB-I00) and the Red Madrileña de Nanomedicina en Imagen Molecular-Comunidad de Madrid (P2022/BMD-7403 RENIM-CM). B.Iz. is supported by the program Atracción de Talento of the Comunidad de Madrid (GN:2022-T1/BMD-23767). J.J.F. is supported by grant RYC-2016-20026, funded by MICIN/AEI/10.13039/501100011033’ and ‘ESF Investing in your future’. The CNIC is supported by the Instituto de Salud Carlos III (ISCIII), the Ministerio de Ciencia, Innovación y Universidades (MICIU) and the Pro CNIC Foundation and is a Severo Ochoa Center of Excellence (grant CEX2020-001041-S funded by MICIN/AEI/10.13039/501100011033). We thank J. Sanchez-González for his contribution to imaging analyses in PESA.
Author information
These authors contributed equally: Miriam Díez-Díez, Beatriz L. Ramos-Neble.
Authors and Affiliations
Centro Nacional de Investigaciones Cardiovasculares, Madrid, Spain
Miriam Díez-Díez, Beatriz L. Ramos-Neble, Jorge de la Barrera, J. C. Silla-Castro, Ana Quintas, Enrique Vázquez, M. Ascensión Rey-Martín, Benedetta Izzi, Lucía Sánchez-García, Inés García-Lunar, Guiomar Mendieta, Virginia Mass, Nuria Gómez-López, Cristina Espadas, Gema González, Antonio J. Quesada, Ana García-Álvarez, Antonio Fernández-Ortiz, Enrique Lara-Pezzi, Ana Dopazo, Fátima Sánchez-Cabo, Borja Ibáñez, Vicente Andrés, Valentín Fuster & José J. Fuster
CIBER de Enfermedades Cardiovasculares, Madrid, Spain
Inés García-Lunar, Ana García-Álvarez, Antonio Fernández-Ortiz, Enrique Lara-Pezzi, Ana Dopazo, Fátima Sánchez-Cabo, Borja Ibáñez, Vicente Andrés & José J. Fuster
Cardiology Department, University Hospital La Moraleja, Madrid, Spain
Inés García-Lunar
Servicio de Cardiología, Institut Clínic Cardiovascular, Hospital Clínic de Barcelona, Barcelona, Spain
Guiomar Mendieta & Ana García-Álvarez
Institut d’Investigacions Biomèdiques August Pi I Sunyer, Barcelona, Spain
Universitat de Barcelona, Barcelona, Spain
Ana García-Álvarez
Hospital Clínico San Carlos, Universidad Complutense, IdISSC, Madrid, Spain
Antonio Fernández-Ortiz
Cardiology Department, IIS-Fundación Jiménez Díaz University Hospital, Madrid, Spain
Borja Ibáñez
Icahn School of Medicine at Mount Sinai, New York, NY, USA
Valentín Fuster
You can also search for this author in PubMed Google Scholar
Contributions
M.D.-D. and B.L.R.-N. analyzed somatic variant data, conducted association studies and contributed to writing the paper. A.Q., E.V., M.A.R-M., N.G-L., C.E. and A.D. conducted DNA sequencing analyses. J.B., J.C.S-C. and L.S-G. performed bioinformatics analyses of sequencing data. F.S.-C. supervised all bioinformatics and statistical analyses. B.Iz. provided methodological input. G.G. and A.J.Q. contributed to biological sample processing and revision. I.G.-L., G.M., V.M., A.G.-A., A.F.-O., E.L.-P., B.Ib., V.A. and V.F. contributed to the phenotyping of the PESA cohort and provided methodological input. V.F. is the principal investigator of the PESA study. J.J.F. conceived this study and its design, supervised all analyses and wrote the paper. All authors provided critical revision of the paper.
Corresponding authors
Correspondence to Valentín Fuster or José J. Fuster .
Ethics declarations
Competing interests.
The authors declare no competing interests
Peer review
Peer review information.
Nature Medicine thanks Alexander Bick, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: Michael Basson, in collaboration with the Nature Medicine team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended data fig. 1 basic features of clonal hematopoiesis (ch)-related mutations and subclinical atherosclerosis burden in the study population..
We performed deep sequencing to identify somatic mutations in a custom panel of 54 CH-related genes in 3,692 middle-aged individuals from the PESA cohort. a , CH-related genes included in the custom panel. Genes for which a CH mutation was found in at least one individual are underlined and in bold font. Genes typically considered as drivers of myeloid CH (frequently known as CHIP, clonal hematopoiesis of indeterminate potential) or lymphoid-CH are shown separately. b , Assay sensitivity for the detection of CH-related variants at different variant allele fractions (VAF). Dashed lines show the median sequencing depth of our study (3,712x; vertical line), and the sensitivity to detect CH variants at the minimum VAF threshold (0.2%) used to define CH (horizontal lines). c , Graphical representation of the CH-related genes found mutated in the study population; font size is proportional to the frequency of mutations. d , Proportions of different type of mutations in CH-related genes according to their functional consequence. e , Proportions of different types of single nucleotide substitutions. f , Distribution of mutant clone size, assessed as VAF, in DNMT3A (n = 657), TET2 (n = 153) and other sequenced genes (n = 362). The dashed line shows the 2% VAF threshold typically used to identify CHIP. Boxes represent the 25th (Q1), 50th (median) and 75th (Q3) percentiles of the data. Whiskers represent Q1 - 1.5*IQR at the minimum and Q3 + 1.5*IQR at the maximum. g , Proportion of CH-related variants identified after downsampling of sequencing depth to 30x, 100x, 400x and 1,000x. “base” indicates the median sequencing depth for these variants using our original sequencing approach (3,625x); n = 168 CH variants, 28 for each VAF range. h , Number of CH-related variants identified after downsampling to specific sequencing depths.
Extended Data Fig. 2 Subclinical atherosclerosis burden at baseline stratified by CH status.
We performed deep targeted sequencing to identify somatic mutations in a custom panel of 54 CH-related genes in 3,692 middle-aged individuals from the PESA cohort. CACS and 3DVUS imaging were used to determine subclinical atherosclerosis burden, assessed cross-sectionally at baseline. Graphs show the proportion of individuals with no detectable atherosclerosis and across tertiles of plaque burden, based on CAC score ( a , n = 3,687), global plaque volume by 3DVUS ( b , n = 3,543), carotid plaque volume by 3DVUS ( c , n = 3,659) or femoral plaque volume by 3DVUS ( d , n = 3,569). Statistical significance for differences in subclinical atherosclerosis burden, defined as 0 or tertiles, was evaluated using an ordinal logistic regression model.
Extended Data Fig. 3 Effects of clonal hematopoiesis (CH) on de novo atherosclerosis development in femoral arteries.
We investigated the association between CH at baseline and de novo development of femoral atherosclerosis ∼ 3-years after enrollment among individuals who initially lacked detectable atherosclerosis in that region, assessed by 3-dimensional-vascular ultrasound (3DVUS) imaging. a , Association between CH with VAF ≥ 2% and de novo atherosclerosis development in femoral arteries, based on multivariate logistic regression analyses adjusted for age, sex, conventional modifiable risk factors as categorical variables and the cumulative systemic burden of atherosclerosis across the timeframe of the study (AUC for CACS, assessed by CT imaging, and global atherosclerotic plaque volumes, assessed by 3DVUS imaging) (n = 1,932). b , Association between CH and de novo atherosclerosis development in femoral arteries, based on multivariate logistic regression analyses adjusted for age, sex, absolute counts of leukocytes, erythrocytes and platelets, and the cumulative systemic burden of atherosclerosis across the timeframe of the study (n = 2,353). c , Association between myeloid CH or CHIP with VAF ≥ 2% and de novo atherosclerosis development in femoral arteries, based on multivariate logistic regression analyses adjusted for age, sex, conventional modifiable risk factors and the cumulative systemic burden of atherosclerosis across the timeframe of the study (n = 1,965). d , Association between CHIP and de novo atherosclerosis development in femoral arteries, based on multivariate logistic regression analyses adjusted for age, sex, absolute counts of leukocytes, erythrocytes and platelets, and the cumulative systemic burden of atherosclerosis across the timeframe of the study (n = 2,353). Bars in each panel indicate 95% confidence intervals centered in the mean value (square).
Extended Data Fig. 4 Representative 3DVUS images from subclinical atherosclerosis burden in femoral arteries across tertiles of atherosclerotic plaque volume.
Multiplanar views (upper rows, scale bar: 2 mm) and 3D reconstruction (lower rows) from the right and left femoral arteries. Images are representative of the upper limits of each tertile of atherosclerotic plaque volume. 3DVUS, 3-dimensional vascular ultrasound.
Supplementary information
Supplementary information.
Supplementary Tables 1, 3–6 and 8, legends for Supplementary Tables 2 and 7 (uploaded in an independent spreadsheet file) and Supplementary Figs. 1–3.
Reporting Summary
41591_2024_3213_moesm3_esm.xlsx.
Supplementary Table 2. CH-related mutations identified in the study population. We performed deep targeted sequencing to identify somatic mutations in a custom panel of 54 CH-related genes in 3,692 individuals from the PESA cohort. A total of 1,172 CH-related somatic variants were identified in 900 participants in PESA at baseline. Ref., reference; Alt., alternate; SNV, single nucleotide variant. Supplementary Table 7. CH-related mutations identified longitudinally in the study population. Using deep targeted sequencing, we found 602 CH variants in 494 individuals from the PESA cohort both at baseline and at the 6-year follow-up. Ref., reference; Alt., alternate; SNV, single nucleotide variant.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/ .
Reprints and permissions
About this article
Cite this article.
Díez-Díez, M., Ramos-Neble, B.L., de la Barrera, J. et al. Unidirectional association of clonal hematopoiesis with atherosclerosis development. Nat Med (2024). https://doi.org/10.1038/s41591-024-03213-1
Download citation
Received : 19 May 2023
Accepted : 25 July 2024
Published : 30 August 2024
DOI : https://doi.org/10.1038/s41591-024-03213-1
Share this article
Anyone you share the following link with will be able to read this content:
Sorry, a shareable link is not currently available for this article.
Provided by the Springer Nature SharedIt content-sharing initiative
Quick links
- Explore articles by subject
- Guide to authors
- Editorial policies
Sign up for the Nature Briefing: Translational Research newsletter — top stories in biotechnology, drug discovery and pharma.
An official website of the United States government
The .gov means it’s official. Federal government websites often end in .gov or .mil. Before sharing sensitive information, make sure you’re on a federal government site.
The site is secure. The https:// ensures that you are connecting to the official website and that any information you provide is encrypted and transmitted securely.
- Publications
- Account settings
Preview improvements coming to the PMC website in October 2024. Learn More or Try it out now .
- Advanced Search
- Journal List
- PMC11343254

This is a preprint.
Evolutionary measures show that recurrence of dcis is distinct from progression to breast cancer, angelo fortunato.
1. Arizona Cancer Evolution Center and Biodesign Center for Biocomputing, Security and Society, Arizona State University, 727 E. Tyler St., Tempe, AZ 85281, USA
2. School of Life Sciences, Arizona State University, 427 East Tyler Mall, Tempe, AZ 85287, USA
Diego Mallo
Luis cisneros, lorraine m. king.
3. Duke University School of Medicine, Durham, NC 27710, USA
4. Department of Medicine, Genetics, and Biomedical Data Science Stanford School of Medicine, Stanford, CA 94305
5. Stanford Cancer Institute, Stanford School of Medicine, Stanford, CA 94305
Christina Curtis
6. Chan Zuckerberg Biohub, San Francisco, CA
Marc D. Ryser
Joseph y. lo, allison hall, jeffrey r. marks, e. shelley hwang, carlo c. maley, associated data.
All the sequencing data used in this manuscript is publicly available. The cross-sectional WES data at SRA with IDs (SRP298346 and XX) and the longitudinal WGS and WES data at HTAN dbGaP’s study accession phs002371.v6.p1.
Progression from pre-cancers like ductal carcinoma in situ (DCIS) to invasive disease (cancer) is driven by somatic evolution and is altered by clinical interventions. We hypothesized that genetic and/or phenotypic intra-tumor heterogeneity would predict clinical outcomes for DCIS since it serves as the substrate for natural selection among cells. We profiled two samples from two geographically distinct foci from each DCIS in both cross-sectional (N = 119) and longitudinal cohorts (N = 224), with whole exome sequencing, low-pass whole genome sequencing, and a panel of immunohistochemical markers. In the longitudinal cohorts, the only statistically significant predictors of time to non-invasive DCIS recurrence were the combination of treatment (lumpectomy only vs mastectomy or lumpectomy with radiation, HR = 12.13, p = 0.003, Wald test with FDR correction), ER status (HR = 0.16 for ER+ compared to ER-, p = 0.0045), and divergence in SNVs between the two samples (HR = 1.33 per 10% divergence, p = 0.018). SNV divergence also distinguished between pure DCIS and DCIS synchronous with invasive disease in the cross-sectional cohort. In contrast, the only statistically significant predictors of time to progression to invasive disease were the combination of the width of the surgical margin (HR = 0.67 per mm, p = 0.043) and the number of mutations that were detectable at high allele frequencies (HR = 1.30 per 10 SNVs, p = 0.02). These results imply that recurrence with DCIS is a clinical and biological process different from invasive progression.
Introduction
The improvement of radiological techniques and preventive screening of breast cancer conducted on a large scale makes it possible to identify mammary gland neoplasms at an early stage of development, when they are still confined within the glandular ducts. This neoplasm is termed ductal carcinoma in situ (DCIS) ( 1 ). Estimates from several natural history studies of DCIS indicate that 20–30% will progress to invasive cancer without definitive surgical treatment ( 2 , 3 ), implying that as many as 70% of patients who have surgery for DCIS may not derive benefit.
The ability to recognize which pre-cancerous tumors are likely to progress to invasive cancer is of great importance because it would identify high-risk patients for surgical, pharmacological, and/or radiation treatment. In contrast, low-risk patients could be managed by watchful waiting, avoiding the unnecessary harms and side effects associated with these therapies ( 4 ). Furthermore, selecting patients most at risk would facilitate reallocating healthcare resources to those who would benefit most from treatment.
Evolutionary mechanisms drive tumor progression ( 5 ). The impairment of control mechanisms of genetic integrity ( 6 ) accelerates the accumulation of new genetic alterations in cancer cells ( 7 ). The combination of these alterations in an increasing number of clones represents a critical factor in tumor progression, as these clones constitute the substrate upon which selection acts ( 8 ). The identification of mutations and the level of genetic (and phenotypic) heterogeneity have been shown to be associated with the risk of tumor progression in other pre-cancers, like Barrett’s esophagus ( 9 – 11 ). The higher the number of mutations and the greater the intratumor genetic heterogeneity, the higher the risk of developing clones that are cancerous, metastatic, and treatment-resistant ( 12 – 16 ).
It is challenging to integrate the combined effect of many mutations and genetic alterations that act simultaneously in cancer cells ( 17 ). Investigating the number of mutations and the level of heterogeneity allows us to introduce a quantitative parameter independent of the functional consequences of specific combinations of mutations, serving as a surrogate measure of the degree of evolvability of the neoplastic cells ( 18 , 19 ).
Both genetic and phenotypic heterogeneity can be measured by comparing different regions of the same tumor, ideally through analysis of longitudinal cohorts with linked clinical outcomes. Such studies often necessitate analysis of archival formalin-fixed paraffin-embedded (FFPE) samples, which is challenging due to partial degradation of the DNA, FFPE-induced artifacts, which manifest as sequence alterations, and low yield of nucleic acids from a limited number of sections. We recently published a workflow that overcomes these challenges, enabling the assessment of measures of genetic divergence between regions of the same tumor ( 20 ). This work aimed to test the hypothesis that genetic and phenotypic heterogeneity within DCIS can predict the recurrence of DCIS and/or progression to invasive ductal carcinoma (IDC).
Materials and Methods
Experimental design.
We performed two observational studies ( Fig. 1 , Table 1 ) to study DCIS progression. In a cross-sectional study ( Fig. 1A ), we compared DCIS samples from patients with DCIS only ( Pure DCIS , n = 58) versus DCIS samples from patients with synchronous invasive ductal carcinoma ( Synchronous DCIS , n = 61). In a longitudinal case-control study ( Fig. 1B ), we collected samples from patients with primary DCIS who were treated and then followed until they were diagnosed with an IDC recurrence ( Progressors , n = 56), were diagnosed with a DCIS-only recurrence ( Recurrents , n = 69), or did not recur within their follow-up time ( Nonrecurrents , n = 99, minimum five years). We calculated the median follow-up time using the reverse Kaplan-Meier method ( 21 ). In both cohorts, we characterized the genotype and phenotype of two DCIS regions per patient. All samples came from different FFPE blocks or were separated by at least 8mm. For some progressors, we also obtained a subsequent IDC sample. The Institutional Review Board (IRB) of Duke University Medical Center approved this study, and a waiver of consent was obtained according to the approved protocol.

A : Cross-sectional study: Synchronous DCIS tumors are presumed to have evolved from pure DCIS that existed before the progression of the synchronous IDC. In patients with synchronous DCIS, only the DCIS component was sampled and assayed unless otherwise specified. B : Longitudinal case-control study: pure-DCIS samples from patients treated and followed up for at least five years or until they progress or recur. n: number of patients per cohort.
Patient Cohorts with WES data, lpWGS, or IHC data.
| Study | Cross-sectional | Longitudinal | |||
|---|---|---|---|---|---|
| Assays | WES/IHC | WES/lpWGS/IHC | |||
| Cohort | |||||
| Number of patients | 58 | 61 | 99 | 69 | 56 |
| Median [min, max] | 2010 [1998, 2015] | 2010.5 [2000, 2017] | 2009 [2000, 2014] | 2008 [1998, 2017] | 2006.5 [1999, 2016] |
| Median [min, max] | 57.2 [34.0, 74.9] | 57.9 [40.9, 74.0] | 55.0 [41.0, 75.0] | 54.9 [40.0, 76.0] | 50.6 [38.0, 73.0] |
| 1 | 1 [1.7%] | 1 [1.6%] | 2 [2.0%] | 6 [8.7%] | 4 [7.1%] |
| 2 | 23 [39.7%] | 22 [36.1%] | 38 [38.4%] | 29 [42.0%] | 17 [30.4%] |
| 3 | 34 [58.6%] | 38 [62.3%] | 59 [59.6%] | 34 [49.3%] | 35 [62.5%] |
| Median [min, max] | 3.3 [0.8, 12.4] | 4.0 [0.2, 10.8] | 2.5 [0.3, 10.2] | 1.7 [0.2, 10.0] | 1.85 [0.4, 14.0] |
| ER (+) | 43 [74.1%] | 40 [65.6%] | 65 [65.7%] | 42 [60.9%] | 27 [48.2%] |
| ER (–) | 8 [13.8%] | 21 [34.4%] | 25 [25.3%] | 17 [24.6%] | 10 [17.9%] |
| ER (unknown) | 7 [12.1%] | 0 | 9 [9.1%] | 10 [14.5%] | 19 [33.9%] |
| Lumpectomy (Lump.) | 5 [5.1%] | 17 [24.6%] | 15 [26.8%] | ||
| Lump. + Radiation (Rad.) | 60 [60.6%] | 44 [63.8%] | 20 [35.7%] | ||
| Mastectomy | 22 [37.9%] | 24 [39.3%] | 33 [33.3%] | 7 [10.1%] | 19 [33.9%] |
| Lump. (Unk. Rad.) | 36 [62.1%] | 37 [60.7%] | 1 [1.0%] | 1 [1.4%] | 2 [3.6%] |
| Median [min, max] | 98 [60, 228] | 37 [12, 196] | 55 [12, 176] | ||
| Ink on tumor | 0 | 0 | 0 | ||
| <2mm | 23 [23.2%] | 28 [40.6%] | 15 [26.8%] | ||
| At least 2mm | 35 [35.4%] | 22 [31.9%] | 20 [35.7%] | ||
| Clear, NA mm | 41 [41.4%] | 19 [27.5%] | 21 [37.5%] | ||
| White | 34 [58.6%] | 43 [70.5%] | 67 [67.7%] | 41 [59.4%] | 28 [50.0%] |
| Black | 22 [37.9%] | 15 [24.6%] | 22 [22.2%] | 21 [30.4%] | 23 [41.1%] |
| Other | 2 [3.4%] | 3 [4.9%] | 2 [2.0%] | 2 [2.9%] | 2 [3.6%] |
| Unknown | 0 | 0 | 8 [8.1%] | 5 [7.2%] | 3 [5.4%] |
Unk. = Unknow.
Clinical specimens
We classified breast tumors according to the World Health Organization (WHO) criteria ( 22 ). We graded the IDC and DCIS samples according to the Nottingham grading system ( 23 ) or recommendations from the Consensus Conference on DCIS classification ( 24 ), respectively.
All samples were obtained from formalin-fixed paraffin-embedded (FFPE) breast tissue blocks. Cases from the cross-sectional studies were obtained from the Duke Pathology archives. Cases from the longitudinal study were obtained from Translational Breast Cancer Research Consortium (TBCRC) sites, a national multi-center consortium of cancer centers that treat breast cancer patients. All cases underwent detailed pathology review (AH) for histologic features and case eligibility.
DNA extraction and sequencing
The DNA extraction, sequencing, and data processing protocol has been previously reported ( 20 ). For each neoplastic sample, we extracted the DNA from multiple serial archival FFPE tissue block sections after macro-dissecting the areas of interest. To estimate the germline sequence, we also extracted DNA from either distant benign breast tissue or a benign lymph node. The study pathologist confirmed the presence of ≥70% neoplastic cells in the microdissected areas of neoplastic samples and their absence from control samples.
After DNA extraction, hybrid capture was performed using two targeted panels (all exons of the 83 genes in the breast cancer gene panel and the human exome), and the multiplexed libraries were sequenced using either an Illumina HiSeq with 4-channel chemistry (cross-sectional study) or a NovaSeq 6000 machine with 2-channel chemistry (longitudinal study). After alignment to the Genome Reference Consortium Human Build 37 and marking duplicates, we obtained a mean de-duplicated depth of 115.9 ± 52.2 (SD). The resulting BAM files were the input data for our SNV calling and heterogeneity calculation pipeline. We discarded samples with less than 40% of the target covered at 40X. Sequencing was performed at the McDonnell Genome Institute at Washington University School of Medicine in St. Louis.
Additionally, we performed low-pass whole genome sequencing for the longitudinal study as previously described ( 25 ). The resulting BAM files were used as the input data for the CNA characterization pipeline.
SNV characterization
We used our previously reported software ITHE ( 20 ) to calculate by-patient SNV burden and divergence, leveraging the two neoplastic geographically distant samples and a control sample from the same patient. We recently developed, optimized, and validated this pipeline using 28 pairs of technical replicates (same extracted DNA, two aliquots were independently sequenced) of macrodissected FFPE DCIS samples similar to the specimens analyzed here. We used the filtering parameters we found optimal previously ( 20 ). ITHE was optimized for accurate divergence estimation and thus tries to maximize variant calling’s precision. We measured SNV divergence as the percentage of mutations detected in the union of the mutations from the two samples that are not shared by both samples. We required that the union set of mutations had at least five mutations to calculate divergence. SNV burden was calculated as the union of mutations in both samples. When comparing DCIS and IDC samples in the cross-sectional study, we report the mean of the two comparisons between one of the two DCIS samples and the IDC sample.
Functional analysis
We performed the functional enrichment analysis of genes that harbored non-synonymous SNV mutations with PANTHER ( 26 ) and DAVID ( 27 , 28 ). We corrected the fold enrichment p-values considering the false discovery rate (FDR).
CNA characterization
We followed our previously published protocols for low-pass WGS data processing and CNA calling ( 25 ). Briefly, we used Nextflow-base’s Sarek pipeline to align the lpWGS data to the GRCh38/hg38 reference genome, marked duplicates, and re-calibrated quality scores. We used the resulting alignments to call autosomal CNA variants using QDNAseq ( 29 ) on 50-kb genomic bins after filtering genomic regions and reads for mappability and QC content while estimating ploidy and purity. We corrected the log2 ratio for the latter. CNAs with ∣ c o r r e c t e d l o g 2 r a t i o ∣ > 0.3 were considered as altered and normal otherwise. To maximize the robustness of our statistics, we measured CNA burden per sample as the proportion of the genome that was altered (over the total genome considered) and CNA divergence per patient as the proportion of the altered genome that is not shared between the two samples over the altered genome per patient (i.e., C N A d i v e r g e n c e = A Δ B A ∪ B , with A and B defined as the set of altered genomic regions of each homonymous sample, and Δ the set symmetric difference operator).
Immunohistochemistry characterization
We chose a series of 15 candidate proteins ( Supplementary Table S1 ) representing several categories including essential breast cancer drivers (ER, PR, HER2), immune-related (FOXP3, CD68), resource and microenvironmental measures (GLUT1, CA9, CD31, FASN), myoepithelial and basement membrane (TP63, COL15A1) and progenitor or stem cell-related (ALDH1 and RANK) markers. Additional proteins included the proliferation marker KI-67, the adhesion marker phospho-FAK, and COX2 (PTGS2), which were previously described as being associated with DCIS progression. In the longitudinal study, these were reduced to ER and GLUT1 only ( Supplementary Tables S2 - 3 ), based on the results from the cross-sectional study and the paucity of samples. We measured stain intensity using detailed expert scoring. In most cases, the study pathologist used a scoring system that captures the distribution of intensities in an IHC profile, while for a smaller number of markers, it was binary ( Supplementary Table S1 ). The IHC profile was quantified as the percentage of the slide presenting different levels of increasing staining intensity: absence, low, medium, and high. Medium staining was deemed as approximately twice as intense as low staining and high staining three times as intense as low staining.
We evaluated the IHC at three different scales of comparison:
- The average intensity of immunofluorescence across samples for each patient, measuring the typical intensity of IHC signal per patient.
- The variance of the intensity between samples for each patient, measuring the variations of IHC signal between distant locations in each patient.
- The variance of intensity within samples, measuring the variations of IHC signal at short distances in each patient.
These three measures are quantified by the Mean of Intensity Score, the Earth Mover’s Distance, and the Cumulative Density Index. Briefly, the Intensity Score is the weighted sum of the IHC profile proportions normalized by the maximum possible staining, the Earth Mover’s Distance represents the minimum cost of turning one profile into another ( 30 ), and the Cumulative Density Index represents how close from a uniform distribution the observed profile is and ranges from 0 (all the profile weight in one of the extreme categories) to 1 (uniform profile). See a detailed description of these statistics in Supplementary Text 1 .
Statistical analysis
Cohort characterization.
For each study, we compared differences in the central tendency of genetic and phenotypic variables per patient between cohorts using the Kruskal-Wallis Rank Sum or the Mann-Whitney U tests for many or two cohorts, respectively. We followed the Kruskal-Wallis Rank Sum test with Dunn’s post-hoc test while controlling for multiple tests using the Holm-Šidák adjustment ( 31 ). Exceptionally, CNA divergence met the assumptions of a parametric test, and thus, we used an ANOVA followed by Tukey HSD post-hoc tests. In cases where we used multiple measurements per patient (CNA burden), we used a Mixed-effects ANOVA with different random effect intercepts per patient to account for data dependencies (on the square-root-transformed variable), followed by Tukey’s HSD on the estimated marginal means.
Distinguishing Pure DCIS from Synchronous DCIS
We performed variable selection among the phenotypic measurements with significant differences between cohorts using a Random Forest classification model ( 32 ) under the Gini impurity criterion to return the importance ranking of each feature given by their predictive power. We used the two top measurements to build a generalized linear logistic model. Similarly, we built a generalized linear logistic model with the genetic measurements that showed significant differences between cohorts and the combination of the three. Due to missing data, we compared the models under the Akaike information criterion (AIC) on the smallest dataset for all models ( 33 ).
Association with clinical outcomes
Using our longitudinal study, we determined whether genetic and phenotypic statistics were independently associated with the time to clinical outcome (non-invasive recurrence or progression) using Cox regression analyses after checking they met the proportional hazards assumption. Nonrecurrent patients were right-censored using their follow-up time, and progressors’ recurrence time was used as their time to clinical outcome. Recurrents were discarded when considering progression, and progressors were discarded when considering non-invasive recurrence. We also provide supplementary results in which the clinical outcomes are “any recurrence” and “progression without discarding recurrent patients.” In this case, recurrents were right-censored at the time of recurrence when considering progression, and otherwise, their recurrence time was used as their time to clinical outcome. We evaluated the statistical significance of Cox regressors using the Wald test. We used the proportional hazard regression model for one variable (SNV burden) to stratify patients into low and high SNV burden and plotted their event-free survival curves. We stratified using the risk relative to the patient with all variables (i.e., SNV burden here) set at the mean value (i.e., type = “risk”, reference = “sample”, in the predict.coxph function of the survival R package). We chose the threshold that maximized Youden’s J statistic ( 34 ) using the true outcomes. In all cases, we used the log-rank test to compare the survival trends of two or more groups.
We also integrated 18 clinical covariates ( Supplementary Table S4 ) with our eight genetic and phenotypic measurements to model time to non-invasive recurrence and time to progression. We performed variable selection using Cox LASSO and chose the regularization parameter that minimized the partial-likelihood deviance via 10-fold cross-validation. To reduce the stochasticity of the results, we performed this process 100 independent times per model and selected the variables that were selected in at least 90% of them. To reduce missingness, we performed mean imputation on the clinical covariates before variable selection. The selected variables were used to build the final Cox regression models using all patients with available (imputed) data for those variables. Alternatively, we selected patients with data for all covariates chosen without imputation. We used the final models to stratify patients as in the univariate proportional hazards regression above. In all cases, the model used to stratify patients and plot their event-free survival curves includes all the variables included in the forest plot. All variables were standardized to make hazard ratios (HRs) comparable, and thus, HRs are relative to a change of 1 standard deviation unless specified otherwise.
Data Availability
Reproducibility.
Scripts to reproduce most data pre-processing and statistical analysis can be found at https://github.com/adamallo/ManuscriptScripts_DCISRecurrenceVsProgression .
Study cohorts
We investigated DCIS progression to invasive cancer using two independent observational studies with different patients: a cross-sectional study and a longitudinal study ( Fig. 1 , Table 1 ). In the cross-sectional study ( Fig. 1A ), we compared DCIS samples from patients with DCIS only ( Pure DCIS , n = 58) versus DCIS samples from patients with synchronous DCIS with invasive ductal carcinoma ( Synchronous DCIS , n = 61). In the separate longitudinal study ( Fig. 1B ), we compared pure DCIS samples from patients who were treated and had long-term follow-up (median = 117 months, 95% CI [104, 132]). This cohort consisted of patients who progressed to IDC ( progressors ) (n = 56), patients who had a DCIS-only recurrence ( recurrents , n = 69), or patients who did not recur during the follow-up interval ( nonrecurrents , n = 99). In both studies, we characterized the genotype and phenotype of two formalin-fixed paraffin-embedded DCIS samples per patient, enabling measures of evolutionary divergence (see Methods ). We also obtained a single sample of their IDC recurrence for some progressors.
Cross-sectional study
Single nucleotide mutational burden.
Pure DCIS carried fewer SNVs per patient (mean 7.5 ± 10.6 standard deviation) than synchronous DCIS (10.4 ± 15.3), but this difference was not statistically significant ( Fig. 2A ).

Distribution of the number of SNVs per patient in the two cross-sectional cohorts A and the two lesion types (DCIS vs. IDC) present in the synchronous cohort B . Distribution of SNV genetic divergence (percentage of private mutations) per patient in the two cross-sectional cohorts C . We calculated divergence for tumors with at least five mutations in the union of the two samples, which explains the lower number of tumors per group. P-values shown if p ≤ 0.1, A, C : Mann-Whitney U, B : Paired-samples sign test. Interquartile range (vertical line) and median (point) in burgundy, N: number of patients.
The invasive component in synchronous DCIS patients showed a statistically significantly increased number of SNVs (18.1 ± 31.5, Fig. 2B ) compared with their DCIS counterpart (Paired-samples sign test, p = 0.04) largely due to four cases of IDC with a dramatic increase in mutation burden.
SNV Genetic Divergence
We measured the SNV genetic divergence as the percentage of mutations that are private to either sample per patient. Synchronous DCIS showed higher genetic divergence (21.5% ± 17.5%) than pure DCIS (10.8% ± 17.4%, Fig. 2C ) (Mann-Whitney U test, p = 0.009). Additionally, we also characterized the genetic divergence between the two synchronous components (i.e., DCIS vs. IDC in synchronous patients) (44.5% ± 29.0%), which is higher than the paired synchronous DCIS divergence ( Supplementary Fig. S1 , Paired-samples sign test, p = 0.002).
Phenotypic characterization
Synchronous DCIS samples presented higher levels of GLUT1 staining ( p = 0.004) and lower levels of CA9 staining ( p = 0.01) than pure DCIS samples ( Fig. 3A , pairwise Mann-Whitney U tests of mean intensity scores [MIS], unadjusted p-values); all other markers showed non-significant differences between groups. This result holds when one of the two DCIS samples per patient is used randomly instead of the MIS ( Supplementary Fig. S2 ).

Distribution of mean intensity scores (MIS) per patient (see Methods ) A , between-sample divergence ( B , Earth Mover’s Distance [EMD]) and within-sample divergence ( C , Cumulative Density Index [CDI]). A: for each patient and IHC marker, B and C: only markers with significant differences between cohorts (unadjusted p-values). Unadjusted pairwise Mann-Whitney U p-values shown if p ≤ 0.1. Interquartile range (vertical line) and median (point) in burgundy. N: number of patients.
Phenotypic Divergence
We characterized the between-sample phenotypic divergence for each marker using a distance between staining intensity profiles (Earth Mover’s Distance) and the within-sample divergence using a measure of staining intensity uniformity (Cumulative Density Index; see Supplementary Methods for detailed definition of these indices).
Multiple markers presented differences in between-sample divergence between pure DCIS and synchronous DCIS samples, with the latter showing increased divergence for GLUT1 ( p = 0.01), FOXP3 ( p = 0.01), and HER2 ( p = 0.04) staining, but decreased divergence of ER ( p = 0.01) staining ( Fig. 3B , Supplementary Fig. S3 , Pairwise Mann-Whitney U tests, unadjusted p-values). This reduction of ER phenotypic divergence in synchronous DCIS samples was replicated in the within-sample measures ( p = 0.01) and mimicked by CA9 ( p = 0.01) ( Fig. 3C , Supplementary Fig. S4 , Pairwise Mann-Whitney U tests, unadjusted p-values). A reduction in the phenotypic divergence for ER in synchronous DCIS samples indicates larger uniformity across and within samples, while the mean intensity of ER signal is not markedly different ( Fig. 3A ).
All eight significant phenotypic divergence features—MIS for GLUT1 and CA9 ( Fig. 3A ), EMD for GLUT1, FOXP3, ER and HER2 ( Fig. 3B ), and CDI for ER and CA9 ( Fig. 3C )—were combined in a mixed logistic regression to model the progression status of the samples, from which the most important features were selected according to their relative predictive power. A reduced logistic model including between-sample diversity (EMD) for GLUT1 and within-sample diversity (CID) for ER had statistically significant coefficients (GLUT1 EMD, p = 0.01; ER CDI, p = 0.01) and spanned 40 pure DCIS cases and 52 synchronous DCIS cases. Therefore, we selected these two IHC markers (GLUT1 and ER) as the targets for phenotypic divergence to be included in the longitudinal study.
Logistic regression showed that the only statistically significant genetic measurement (SNV divergence) was strongly associated with the cohort, with p = 0.0136, so it was also selected for evaluation in the longitudinal study.
Longitudinal Study: Associations with Recurrence and Progression
We used the cross-sectional cohort as a discovery cohort, using the synchronous DCIS as a proxy for high-risk DCIS likely to progress to IDC. Samples in our validation cohorts come from patients with pure DCIS with known outcomes ( nonrecurrent , recurred as DCIS, progressed to IDC) and were obtained before treatment ( Fig. 1B ). We sequenced the exomes of two regions of each index DCIS in the longitudinal cohorts, mirroring the methods for the cross-sectional cohorts, and also performed low-pass whole genome sequencing data for most samples.
Mutational burden
Primary DCIS tissue from nonrecurrent patients carried the fewest SNVs (13.4 ± 18.2), followed by that of recurrent patients (19.2 ± 26.4) and progressors (39.7 ± 46.2). These relationships between cohorts were mirrored by the CNA alteration burden ( nonrecurrents : 15.9% ± 15.0% genome altered, recurrents : 17.3% ± 14.8%, progressors : 24.6% ± 17.1%) but presented higher p-values. Thus, SNV burden shows statistically significant differences between nonrecurrents and progressors ( p = 0.003) and between recurrents and progressors ( p = 0.05, Dunn’s test corrected for multiple tests with the Holm-Šidák adjustment) ( Fig. 4A ). In contrast, CNA burden was significantly different only between nonrecurrents and progressors ( p = 0.03, Tukey HSD) ( Fig. 4B ).

Distribution of SNV ( A , C ) and CNA ( B , D ) mutational burdens ( A , B ) and divergences ( C , D ) in the three longitudinal cohorts (Nonrec: nonrecurrents , Rec: recurrents , Prog: progressors ). A : number of SNVs per patient; Omnibus test: Kruskal-Wallis Rank Sum, Post-hoc test: Dunn’s test with control for multiple tests using the Holm-Šidák adjustment. B : proportion of genome with copy number alterations per sample; Omnibus test: Mixed-effects ANOVA on the square-root-transformed proportion of genome altered, Post-hoc test: Tukey HSD on estimated marginal means. C : percentage of private SNV mutations per patient; Omnibus test: Kruskal-Wallis Rank Sum. D : percentage of the genome with copy number alterations private to either sample per patient; Omnibus test: ANOVA, Post-hoc test: Tukey HSD. P-values shown if adjusted p ≤ 0.1. Interquartile range (vertical line) and median (point) in burgundy, N: number of data points ( A , C , and D : patients, B : samples). We only calculated divergence for tumors with at least five mutations in the union of the two samples, which explains the lower number of tumors in C .
Genetic Divergence
Similar to SNV divergence, we measured CNA divergence as the percentage of the altered genome that is private to either sample per patient. SNV divergence was highest in recurrent patients but not statistically different between cohorts ( nonrecurrents : 17.0% ± 13.8%, recurrents : 28.2% ± 25.5%, progressors : 18.4% ± 19.3%, Fig. 4C ). In contrast, CNA divergence followed a decreasing pattern of divergence with progression ( Fig. 4D ), by which nonrecurrents were the most divergent (77.4% ± 16.4%), followed by recurrents (67.7% ± 23.4%) and progressors (63.7% ± 21.7%). Only progressors and nonrecurrents showed statistically significant differences in CNA divergence in pairwise comparisons ( Fig. 7B , p = 0.03, Tukey HSD).

Forest plots describing proportional hazard regressions using variables selected with LASSO ( A , C ) and corresponding Kaplan-Meier plots of patients stratified by the relative risk threshold that maximizes Youden’s J statistic of the outcomes ( B , D ). A-B : Non-invasive-recurrence-free survival. C-D : Progression-free survival. Hazard Ratios (second column, A , C ) are relative to 1 standard deviation. Lumpectomy Only is compared to Lumpectomy + Radiation and Mastectomy and ER+ is compared to ER-. No microcalc(ification)s is compared to having microcalcifications in DCIS-only and/or benign ducts. Tables below Kaplan-Meier plots show the number of samples at risk at different times. Log-rank test.
Functional analysis of non-synonymous SNV mutations
The functional analyses highlighted significant differences between the three cohorts. According to DAVID , recurrent patients showed enrichment of mutated genes involved in taste reception (TAS2R30, TAS2R31, TAS2R43, and TAS2R46), while progressors showed enrichment of genes typically mutated in cancers such as endometrial, small cell lung, prostate, and breast cancer, glioma and melanoma (PIK3CA, ERBB2, PTEN, AKT1, PIK3R2, TP53, PIK3CG), and genes involved in the determination of cell shape, arrangement of transmembrane proteins, and organization of organelles (SPTA1, SPTBN5, DST, SPTAN1). Nonrecurrents did not show significant functional enrichment ( Supplementary Table S5 ). In addition, PANTHER functional analysis revealed an enrichment of several pathways only in progressors ( Supplementary Table S6 ), such as Hypoxia response via HIF activation ( p < 0.001, false discovery rate correction herein this section), Insulin/IGF pathway-protein kinase B signaling cascade ( p < 0.001), p53 pathway ( p = 0.003), Endothelin signaling pathway ( p = 0.003), Hedgehog signaling pathway ( p = 0.02), and PI3 kinase pathway ( p = 0.03).
Phenotypic Characterization and Divergence
We characterized the DCIS phenotypes of the three cohorts using the immunohistochemical profiles of the two markers that showed the highest discriminating power between the two cross-sectional cohorts, ER and GLUT1 (within-sample and between-sample divergence, respectively; see Immunohistochemistry characterization methods section). GLUT1 intensity was different between longitudinal cohorts ( Fig. 5A , p = 0.04, Kruskal-Wallis Rank Sum), like in the cross-sectional study ( Fig. 3A ), with progressors having a generally higher intensity than nonprogressor cohorts, but the pairwise differences were not statistically significant (vs. nonrecurrents p = 0.06, vs. recurrents p = 0.07). ER intensity ( Fig. 5B ) was higher in ER+ progressors ( p = 0.02) and recurrents ( p = 0.03) than in nonrecurrents (Dunn’s test corrected for multiple tests with the Holm-Šidák adjustment). This new pattern was not found in the cross-sectional study, and the difference between progressors and nonrecurrents is robust to ER status stratification ( Supplementary Fig. S5 ).

Distribution of mean normalized intensities (MIS) per patient (see Methods ) in the three longitudinal cohorts (Nonrec: nonrecurrents , Rec: recurrents , Prog: progressors ). A : GLUT1 marker, B : ER marker in ER+ patients only. Omnibus test: Kruskal-Wallis Rank Sum, Post-hoc test: Dunn’s test with control for multiple tests using the Holm-Šidák adjustment. P-values shown if adjusted p ≤ 0.1. Interquartile range (vertical line) and median (point) in burgundy. N: number of patients.
We assessed the phenotypic divergence for these two markers using the same methodology as in the cross-sectional study, evaluating ER within-sample divergence and GLUT1 between-sample divergence, but neither showed a statistically significant difference between longitudinal cohorts ( Supplementary Fig. S6 ).
We tested if our genetic and phenotypic markers were independently associated with the time to non-invasive recurrence or progression using Cox regression analyses. Additionally, alternative clinical outcomes (recurrence [including progression] and progression with non-invasive recurrents right-censored) can be found in the supplementary materials ( Supplementary Figs. S7 - S8 , S10 , Supplementary Tables S7 - S8 ).
Time to non-invasive recurrence was associated with divergences: SNV ( p = 0.024), within-sample ER ( p = 0.026), and CNA ( p = 0.038), while time to progression was primarily associated with totals: SNV burden ( p < 0.0001), ER intensity ( p = 0.025), GLUT1 intensity ( p = 0.027), and CNA burden ( p = 0.045), but also CNA divergence ( p = 0.025) ( Supplementary Tables S9 - S10 , Wald test). The association between SNV burden and progression was the only one that survived multiple-test correction ( Supplementary Tables S7 - S8 , progression adjusted p < 0.0001, Holm correction). Accordingly, we show the capability of this genetic measurement to stratify patients’ non-invasive-recurrence-free ( Fig. 6A ) and progression-free ( Fig. 6B ) survival by splitting patients into low and high SNV burden categories and comparing their event-free survival curves. The Kaplan-Meier plots show differences in the event-free survival curves, with median times to event that differ between groups 100 months for non-invasive recurrence ( Fig. 6A , p = 0.026) and 57 months for progression ( Fig 6B , p < 0.0001, Log-rank test).

Kaplan-Meier plots of stratified patients. A : Non-invasive-recurrence-free survival. B : Progression-free survival. SNV burden thresholds maximize Youden’s J statistic of the outcomes (17 SNVs for non-invasive recurrence and 21 for progression). Log-rank test. The table below the Kaplan-Meier plot shows the number of samples at risk at different times.
Finally, we integrated 18 clinical covariates ( Supplementary Table S4 ) with our genetic and phenotypic measurements to develop comprehensive models of DCIS non-invasive recurrence and progression. Proportional hazard regressions built with variables selected using LASSO contained three significant variables for non-invasive recurrence ( Fig. 7A , treatment option p < 0.001, ER status p = 0.003, and SNV divergence p = 0.018, Wald test) and two for progression ( Fig. 7C , surgical margin p = 0.017, and SNV burden p = 0.004, Wald test), and event-free survival curves of patients stratified using their relative risk were highly significant, with median time to events that differ between groups in 123 months for non-invasive recurrence ( Fig. 7B ) and > 69 months for progression ( Fig. 7D ). An alternative parameterization of the surgical margin as a 2mm threshold showed very similar results ( Supplementary Fig. S9 , p = 0.048, Wald test). The associations with the treatment option and ER status were repeatable without using covariate imputation ( Supplementary Fig. S10 ), while the surgical margin association was only robust when not excluding recurrent patients ( Supplementary Figs. S11 - S12 ). No other significant variables in these models were imputed.
Evolutionary measurements summarize the results of complex evolutionary dynamics, and equivalent observations may result from very different evolutionary scenarios. For example, both a low mutation rate under neutral evolution and a hard selective sweep can generate low divergence of high allele-frequency mutations. Divergence also has multiple scales, and multiple evolutionary processes may affect scales differently or even in opposite directions. Clonal expansion may reduce within-sample divergence but increase between-sample divergence. Intra-tumor heterogeneity provides the fuel for natural selection, but it is not clear what form of intra-tumor heterogeneity (genetic, epigenetic, or phenotypic) is most relevant to the clinical outcomes of a particular tumor, and it is not clear how best to measure it ( 18 ).
The improved efficacy of preventive screenings provided the ability to identify many tumors in the earliest phases of their evolution, demanding the development of new approaches to stratify the risk to these patients to avoid over- and undertreatment. However, every neoplasm develops a unique set of alterations through somatic evolution ( 18 ), making it unlikely that any given set of molecular markers will be universally applicable, even within a given cancer type. In contrast, measures of the evolvability of a neoplasm, such as the number of mutations and measures of intra-tumor heterogeneity, may be universal biomarkers that predict neoplastic progression in many different types of cancers and pre-cancers ( 9 , 10 , 15 ). By taking two spatially distinct samples for each primary pre-cancer, we measured genetic and phenotypic divergence within and between samples, and their relationship with two key clinical processes: 1) recurrence of precancer following treatment and 2) progression of precancer to invasive cancer.
DCIS recurrence and progression are different biological processes
Based on our results, progression from DCIS to invasive breast cancer appears to be a qualitatively and biologically different process from recurrence of DCIS. We had assumed that progression to invasion first requires recurrence of the DCIS and so expected that the factors that predicted recurrence would also predict progression. We were surprised that there was no overlap in their multivariate models ( Fig. 7 ).
Among all genetic and phenotypic variables, SNV burden, as measured with our previously released software ITHE ( 20 ), was the variable that showed the largest differences between the patients that did not recur, the patients that recurred with DCIS, and the patients that progressed to IDC. SNV burden also had the strongest independent association with time to progression and was an essential component of its best multivariate model. The lpWGS CNA burden from the same samples corroborated this finding with higher p-values. Theoretically, this increase in mutation burden may result from an increase in mutation rate, evolutionary time, or self-renewing cell population size. However, due to limitations in detecting variants at low allele frequency, measured mutation burdens are biased towards high allele frequency mutations and are thus most sensitive to early increases in mutation rates or the selective evolutionary forces that drive clonal expansion ( 35 , 36 ). This bias is especially true when using our program ITHE since, by design, it maximizes specificity in exchange for a lower sensitivity for low-frequency mutations in a sample. For these reasons, we do not necessarily expect SNV burden measured differently to show the associations found here.
Progression was also associated with two other magnitude measurements (i.e., totals: ER and GLUT1 intensities) but did not provide enough additional information over the SNV burden to be included as significant variables in the best multivariate model, which also included the size of the surgical margin as a significant predictor.
Previous studies have shown that surgical margins are clinically important in reducing the risk of ipsilateral breast tumor recurrence after breast-conserving surgery ( 37 , 38 ). Positive margins (i.e., DCIS at the edge of the resected tissue) clearly increase recurrence risk, but patients with positive margins were excluded from our study. Instead, we analyzed how the size of the negative margins associate with the clinical outcome. The evidence for this association is mixed in the literature ( 37 , 39 ), but current consensus guidelines consider margins >2mm adequate. Notably, these studies do not typically differentiate recurrence of DCIS from progression to invasive disease in their endpoints, as we did here. We found that the size of the surgical margins was one of the strongest predictors of progression but was not a statistically significant predictor of recurrence with DCIS, neither in the selected multivariate model nor in isolation. This negative result may be due to a type II error, but even if such an association exists, it is likely to be weaker than that observed for progression. We hypothesize that a micro-invasive phenotype could reduce the probability of obtaining large surgical margins, or a phenotype that makes DCIS cells more independent could allow small clusters of cells left over during surgical treatment to survive and further progress to invasive disease more readily. This finding highlights the importance of segregating non-invasive recurrence from progression and how associations with recurrence (of any kind) are primarily a composite of the associations with non-invasive recurrence and progression ( Supplementary Fig. S8A ). We confirmed our results using the consensus guideline >2mm threshold instead of treating surgical margins as a continuous variable, obtaining equivalent though weaker results. This observation shows prognostic information in the size of the surgical margin. The fact that all associations with progression held independently of whether we excluded recurrent patients or right-censored them at the time of DCIS recurrence ( Supplementary Fig. S8C - D ) shows their robustness and adds evidence towards non-invasive recurrence and progression being qualitatively different phenomena.
In contrast, time to non-invasive recurrence was associated with the extent of genetic divergence of SNVs between the two assayed regions of DCIS. We could not corroborate this finding with CNA divergence, which followed the opposite trend but was also correlated to time to recurrence in the univariate models. The true (i.e., known without error) amount of genetic divergence measured using different mutation types should yield equivalent results if large enough mutational burdens of both types are accumulated. A few estimation biases may explain the discordance we observed between SNV and CNA divergences. A low CNA burden may increase the estimated divergence due to a higher false positive rate in the segmentation process without a broad range of true relative intensity values. In fact, CNA burden and CNA divergence were moderately anticorrelated across the study (ρ = −0.36, p < 0.001), and this anticorrelation was driven by the cohort with the lowest CNA burden. High within-sample heterogeneity is also expected to reduce the accuracy of between-sample divergence estimates and lead to the underestimation of the mutation burden. Low SNV burden also leads to missing data in SNV divergence estimates since divergence cannot be calculated accurately with few alterations. ER divergence followed the same direction as CNA divergence, with greater divergence associated with a lower risk of recurrence, but SNV divergence followed the opposite trend. These divergences were the only three measurements associated with time to non-invasive recurrence in the univariate analyses ( Supplementary Table S9 ). Non-invasive recurrence is associated exclusively with divergence statistics, while progression was primarily associated with totals (SNV burden and mean GLUT1 intensity). Intratumor heterogeneity can arise from an increase in the amount of evolution (same mechanisms as mutation burden above) but also with diversifying selection, and we have previously associated it with poor prognosis in other pre-cancers ( 9 ).
Non-invasive recurrence was also associated with the type of DCIS treatment and estrogen-negative status. The fact that patients treated with lumpectomy alone were more likely to recur than those treated with lumpectomy and radiation or mastectomy has been well described. Adjuvant radiation therapy has been previously shown to reduce the risk of recurrence ( 40 ), and after mastectomy, patients are no longer screened using mammograms, making it unlikely that asymptomatic noninvasive recurrences would be detected. The association between recurrence and ER status may be unsurprising since patients with ER+ breast cancers have better prognoses than ER- ones ( 41 , 42 ). However, its association with DCIS recurrence is unclear ( 43 – 45 ), and the balance of evidence points against it ( 46 ). As for surgical margins, most studies are limited by not differentiating between recurrence and progression endpoints. At least one of the studies that made this distinction ( 43 ) found a decrease in non-invasive, but not in invasive recurrences in ER+ patients, which is consistent with our results. Different endpoints may partially explain the mixed evidence on the association between ER and DCIS recurrence and progression.
Functional genetic analysis also showed a difference between the three cohorts, particularly between those DCIS that recurred compared to those that progressed. DCIS that will recur without invasion shows enrichment of mutations in genes involved in the TAS2R signaling network. The activation of these genes determines a pro-apoptotic, anti-proliferative, and anti-migratory response action in highly metastatic breast cancer cell lines ( 47 ). These genes also appear to be involved in the regulation of apoptosis in head and neck squamous cell carcinoma, and their impairment could favor the survival of cancer cells ( 48 ). On the other hand, DCIS that will progress to invasion demonstrates a broader variety of biological processes and pathways involved, such as hypoxia response, insulin/IGF, endothelin, hedgehog, p53, and PI3 kinase signaling pathways. These biological processes are typically altered in various types of cancer and also show an enrichment of mutations in genes involved in the reorganization of the cytoskeleton. The ability to metastasize outside the mammary gland and to relapse observed in these patients is supported by mutations in those pathways.
Synchronous DCIS is not a good model for DCIS progression
Cross-sectional studies are much less resource-intensive, faster, and simpler to conduct than longitudinal cohort studies. If synchronous DCIS (adjacent to IDC) was a good model for primary DCIS that later progressed to IDC, cross-sectional studies could be more readily employed as relevant surrogates for cancer progression. However, our results show this is not possible for our purpose, and in fact, synchronous DCIS shares more similarities with DCIS that will recur as DCIS than with DCIS that will progress.
The pure DCIS samples in our cross-sectional study are equivalent to a mixture of samples from the three cohorts in our longitudinal study since their future outcomes are not considered. Thus, characteristics associated with clinical outcomes are expected to be mixed in the cross-sectional study. We found that DCIS adjacent to IDC showed increased divergence, which may result from divergent evolution facilitated by longer evolutionary times, the interaction with IDC, or an intrinsic characteristic of early-progression DCIS. If we assume that IDC originates from DCIS (stepwise progression model), synchronous DCIS samples are (on average) evolutionarily older than pure DCIS samples, representing a later evolutionary stage than samples from either study. In this case, the cross-sectional study would reveal differences between early and late DCIS. Alternatively, if we assume that an early progression model is also possible (i.e., born to be bad ( 49 )), synchronous DCIS would be enriched with this DCIS sub-type. In this case, the cross-sectional study would show evolutionary characteristics that distinguish those DCIS fated for invasive progression. Additionally, the presence of IDC near synchronous DCIS may also alter its characteristics, modifying its environment systemically (e.g., immune response) and locally (e.g., microenvironment and cell composition through cell migration).
The higher between-sample genetic divergence we found in synchronous DCIS compared to pure DCIS aligns better with stepwise DCIS progression, in which late DCIS would have had more evolutionary time to undergo divergent evolution. Under the early progression model, this may be an intrinsic characteristic of such a DCIS subtype that could facilitate the rapid invasion of nearby tissues. Most (75%) markers with significantly different between-sample divergences showed higher divergence in synchronous DCIS, and all markers with significantly different within-sample divergences showed the opposite trend. These results are concordant with the genetic results and our expectations under a stepwise progression model but did not survive multiple-test correction.
Integrating the results with clonal evolution in neoplastic progression
The two observational studies we conducted here are complementary and together improve our understanding of the evolutionary process leading to DCIS progression and recurrence. We find that primary DCIS that will progress to IDC is more genetically and phenotypically evolved, with higher SNV and CNA burden and more aggressive phenotypes, both metabolically and with respect to its estrogen sensitivity. At least one selective sweep is likely a part of their evolutionary history, which would reduce genetic divergence in the tumor. Higher cell motility could also reduce between-sample heterogeneity. Surgical margins show the strongest association with progression, suggesting that there may be features of the growth pattern of these lesions that make it more difficult to completely excise surgically. In contrast, DCIS recurrence may be primarily enabled by suboptimal clinical management. The few evolutionary features associated with DCIS recurrence suggest an increased accumulation of evolutionary changes in those lesions compared to those that do not recur, which nevertheless do not attain the degree of divergence necessary for invasive progression. In aggregate, the evolutionary history of DCIS recurrences may lack the strong selective sweeps that may be necessary conditions to invade other tissues successfully. DCIS adjacent to IDC shows increased divergence, which may result from divergent evolution facilitated by longer evolutionary times, the interaction with IDC, or an intrinsic characteristic of early-progression DCIS (i.e., born to be bad).
Conclusions
In summary, the evolutionary and clinical measures that predict the recurrence of DCIS differ from those that predict progression to IDC. Furthermore, DCIS adjacent to concurrent invasive cancer appears to be distinct from DCIS that will progress to invasive cancer over time. These findings suggest that the biological dynamics that make DCIS likely to recur differ from those that make it likely to progress, and those dynamics interact differently with our clinical interventions. These insights have the potential to improve both risk stratification and individualized patient management for high-risk DCIS.
Significance
Evolutionary measures of breast pre-cancers associate with local recurrence after surgery, as well as progression to cancer. Recurrence and progression are different biological processes impacted differently by clinical interventions.
Supplementary Material
Supplement 1, acknowledgments.
We thank the Research Computing at Arizona State University for providing HPC ( 50 ) and storage resources that have contributed to the research results reported here. This work is supported in part by NIH grants U54 CA217376, U2C CA233254, R21 CA257980, and R01 CA140657, as well as CDMRP Breast Cancer Research Program Award BC132057 and the Arizona Biomedical Research Commission grant ADHS18-198847. The findings, opinions, and recommendations expressed here are those of the authors and not necessarily those of the universities where the research was performed or the National Institutes of Health.
Conflict of Interest Statement: The authors declare no potential conflicts of interest
- Alzheimer's disease & dementia
- Arthritis & Rheumatism
- Attention deficit disorders
- Autism spectrum disorders
- Biomedical technology
- Diseases, Conditions, Syndromes
- Endocrinology & Metabolism
- Gastroenterology
- Gerontology & Geriatrics
- Health informatics
- Inflammatory disorders
- Medical economics
- Medical research
- Medications
- Neuroscience
- Obstetrics & gynaecology
- Oncology & Cancer
- Ophthalmology
- Overweight & Obesity
- Parkinson's & Movement disorders
- Psychology & Psychiatry
- Radiology & Imaging
- Sleep disorders
- Sports medicine & Kinesiology
- Vaccination
- Breast cancer
- Cardiovascular disease
- Chronic obstructive pulmonary disease
- Colon cancer
- Coronary artery disease
- Heart attack
- Heart disease
- High blood pressure
- Kidney disease
- Lung cancer
- Multiple sclerosis
- Myocardial infarction
- Ovarian cancer
- Post traumatic stress disorder
- Rheumatoid arthritis
- Schizophrenia
- Skin cancer
- Type 2 diabetes
- Full List »
share this!
August 27, 2024
This article has been reviewed according to Science X's editorial process and policies . Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
trusted source
'Low-intensity' blood stem cell transplants for sickle cell appear safe for lung health
by National Institutes of Health
So-called low-intensity blood stem cell transplants, which use milder conditioning agents than standard stem cell transplants, do not appear to damage the lungs and may help improve lung function in some patients with sickle cell disease (SCD), according to a three-year study of adults who underwent the procedure at the National Institutes of Health (NIH).
Damage to lung tissue and worsened lung function is a major complication and leading cause of death in people with sickle cell disease, a debilitating blood disorder. The new study, published today in the Annals of the American Thoracic Society , helps answer whether less intensive types of transplants, which tend to be better tolerated by many adults, by themselves either cause or promote further harm to the lungs.
"By using a low-intensity blood stem cell transplant for sickle cell disease, we may be able to stop the cycle of lung injury and prevent continued damage," said study lead Parker Ruhl, M.D., an associate research physician and pulmonologist at NIH.
"Without the ongoing injury, it's possible that healing of lung tissue might occur, and this finding should help reassure adults living with sickle cell disease who are considering whether to have a low-intensity stem cell transplant procedure that their lung health will not be compromised by the transplant."
Until recently, bone marrow and blood stem cell transplants were the only cure for sickle cell disease, but relatively few adults have undergone the treatments due to health risks associated with high doses of chemotherapy required to prepare for transplants. In addition, the process requires a genetically well-matched donor, usually a sibling who does not have SCD.
These procedures involve giving patients blood stem cells obtained from a donor to grow normal red blood cells to replace the "sickled" cells. The sickled cells block blood flow throughout the body, causing a host of problems, including episodes of acute pain, infections, stroke, and acute chest syndrome, in which lungs are deprived of oxygen.
Researchers say at least one-third of the sickle cell stem cell transplants performed are low-intensity. While they are slightly less effective than the standard transplants, adults who often have more pre-existing organ damage than children tend to do better with them and also experience a lower risk for complications such as graft-versus-host disease . The current study examined if these transplants offered other benefits for adults with already vulnerable lungs.
For the research, Ruhl and her team studied 97 patients with sickle cell disease who underwent a low-intensity, or non-myeloablative, blood stem cell transplant between 2004–2019 at the NIH's Clinical Center in Bethesda, Maryland. Participants were then followed for up to three years.
The researchers conducted a variety of pulmonary function tests, including forced expiratory volume in one second (FEV-1), which measures the amount of air exhaled in the first second after forced exhalation. Another was a lung diffusion test, or diffusing capacity of the lungs for carbon monoxide (DLCO), which measures how much oxygen moves from the lungs to the blood when exhaling. They also conducted a six-minute walk distance test, which measured how far a patient could walk and their oxygen levels during a set time.
After three years, overall lung function among the patients remained stable. FEV-1 levels remained relatively unchanged post-transplant compared to pre-transplant, indicating that lung function did not worsen over time. Notably, DLCO levels and six-minute walk distance improved significantly following transplant.
Ruhl said that larger studies with longer follow-up periods and the inclusion of transplant data from other clinical centers, including those from patients who received a standard transplant, are still needed to put the current findings in context. In the meantime, she and her team will continue to follow the NIH patients and report on longer term outcomes at the five- and 10-year mark.
In December 2023, the U.S. Food and Drug Administration approved two genetic therapies that use patients' own blood stem cells to treat SCD. Researchers hope that the techniques used in this study will also be used to evaluate lung function for other new genetic therapies.
Explore further
Feedback to editors

Not just a 'bad guy': Researchers discover neuroprotective function of Tau protein
5 hours ago

Study shows how common genetic variants in Black Americans increase Alzheimer's risk
6 hours ago

Silicon exoskeletons for blood cells: Engineered blood cells successfully transfused between species
7 hours ago

Neuroscientists explore the intersection of music and memory

Scientists discover a new cardiovascular risk factor and identify a drug able to reduce its effects
8 hours ago

Norovirus study shows how bile acids in breast milk affect newborn gut health

Immunotherapy research offers new hope for children with primary liver carcinomas
9 hours ago

Study unveils novel treatment for blocking SARS-CoV-2 entry into cells

Novel pathway could lead to potential treatment for metabolic liver disease

Music study links dulled emotional reactions in anhedonia with prolonged activity of the brain's attentional networks
10 hours ago
Related Stories

Novel gene-editing therapy continues to show positive results in sickle cell patients
Jun 15, 2024
Researchers suggest B-cells may play role in lung transplant rejection
May 7, 2024
New small molecule could treat sickle cell disease in adults that don't respond to hydroxyurea, alone
Jul 31, 2024
Bone marrow transplant may halt brain blood vessel disease in adults with sickle cell disease
Feb 2, 2023

Studying the impact of stem cell transplants
Mar 9, 2023

'Incompatible' donor stem cells cure adult sickle cell patients
Apr 25, 2018
Recommended for you

Blocking the longevity gene S6K1 extends lifespan in mice by reducing inflammation
Aug 29, 2024

Immune protection against tuberculosis reinfection driven by cells that dampen lung inflammation, shows study
Let us know if there is a problem with our content.
Use this form if you have come across a typo, inaccuracy or would like to send an edit request for the content on this page. For general inquiries, please use our contact form . For general feedback, use the public comments section below (please adhere to guidelines ).
Please select the most appropriate category to facilitate processing of your request
Thank you for taking time to provide your feedback to the editors.
Your feedback is important to us. However, we do not guarantee individual replies due to the high volume of messages.
E-mail the story
Your email address is used only to let the recipient know who sent the email. Neither your address nor the recipient's address will be used for any other purpose. The information you enter will appear in your e-mail message and is not retained by Medical Xpress in any form.
Newsletter sign up
Get weekly and/or daily updates delivered to your inbox. You can unsubscribe at any time and we'll never share your details to third parties.
More information Privacy policy
Donate and enjoy an ad-free experience
We keep our content available to everyone. Consider supporting Science X's mission by getting a premium account.
E-mail newsletter

IMAGES
VIDEO
COMMENTS
This study used a systematic review and meta-analysis as a method to investigate whether STEM enactment in Asia effectively enhances students' learning outcomes. Verifiable examples of science, technology, engineering, and mathematics (STEM) education, effectively being applied in Asia, are presented in this study.
Social, motivational, and instructional factors impact students' outcomes in STEM learning and their career paths. Based on prior research and expectancy-value theory, the study further explored how multiple factors affect students in the context of integrated STEM learning. High school STEM teachers participated in summer professional development and taught integrated STEM to students ...
With the rapid increase in the number of scholarly publications on STEM education in recent years, reviews of the status and trends in STEM education research internationally support the development of the field. For this review, we conducted a systematic analysis of 798 articles in STEM education published between 2000 and the end of 2018 in 36 journals to get an overview about developments ...
For this reason, a meta-analysis study on this subject is considered necessary to investigate the quantitative results of existing studies which have examined the effect of STEM education on academic achievement. The current study set out to synthesize these results and establish the overall magnitude of the effect.
Abstract and Figures. This qualitative descriptive research explored the perspectives of STEM (science, technology, engineering, and mathematics) senior high school students in a public secondary ...
The paper found out that senior high school students are generally interested in the field related to biology. The alignment to the preferred course in college is the primary reason of the participants for enrolling in STEM. Almost all the students wanted to pursue STEM-related careers after their university graduation.
Abstract This study draws upon social cognitive career theory and higher education literature to test a conceptual framework for understanding the entrance into science, technology, engineering, and mathematics (STEM) majors by recent high school graduates attending 4-year institutions. Results suggest that choosing a STEM major is directly influenced by intent to major in STEM, high school ...
The majority of the 127 projects focused on individual STEM disciplines, especially mathematics. The findings, based on IES-funded projects, provided a glimpse of the research input and trends in STEM education in the USA, with possible implications for developing STEM education research in other education systems around the world.
Other research, however, indicates that college environments do impact success for STEM students. Ost (2010), in his study at a large research university, found that positive peer influences can increase the probability of persistence. Although Ost found a large persistence gap between European American students and students of color, he also discovered that students most unlikely to persist ...
This study explored research trends in science, technology, engineering, and mathematics (STEM) education. Descriptive analysis and co-word analysis were used to examine articles published in Social Science Citation Index journals from 2011 to 2020. From a search of the Web of Science database, a total of 761 articles were selected as target samples for analysis. A growing number of STEM ...
It also contributed to the development of 21st-century abilities for all learners. This study determined the status of STEM students in the Philippines. Through descriptive research, this study described and analyzed the trends in the enrolment of STEM students as well as the factors that explain the dropout rate of STEM students.
This qualitative study employed a case research design which sought to investigate nature of the challenges in STEM learning among senior high school students in the Philippines.
Abstract STEM education is getting popular in all over the countries. In order to succeed in this new information-based and highly technological society, students need to develop their ...
This descriptive quantitative study aims to determine why by examining the specific factors 243 12th-grade Saint Louis University Laboratory High School - Senior High (SLU LHS-SH) STEM students considered before selecting the strand, their level of satisfaction with the strand based on these factors, and their Career Aims.
This study employed a narrative-qualitative research design to explore the reasons and experiences of college students who are currently enrolled at a university in San Pablo City and have experienced course mismatch. Three (3) former STEM students qualified as participants via purposive sampling were selected in order to extract relevant and ...
ABSTRACT Interest in science, technology, engineering and mathematics (STEM) has lately received attention in research due to a gap between the number of STEM students and the needs of the labour market. As interest seems to be one of the most important factors in deciding what to study, we focus in the present study on how STEM-interested students weigh multiple interests in making ...
Why do senior high school students choose STEM careers? This study explores their perspectives, motivations, and challenges in pursuing their academic and professional goals.
The study of STEM subjects, she continued, teaches critical-thinking skills, and instills a mindset that will help students find success across numerous areas and disciplines.
ABSTRACT In the study, a meta-analysis study was conducted in order to determine the effects of STEM educational practices on the learning outcomes of the students in the education-training process. The articles were made in national and international context with statistical data that could be included in the meta-analysis study in accordance with the research problem between 2010-2017 ...
Despite reforms, policies, and initiatives implemented worldwide to increase STEM students' interest, attrition is a problem that beset Science education and STEM-related workforce.
Here are 10 qualitative research topics for STEM students: Exploring the experiences of female STEM students in overcoming gender bias in academia. Understanding the perceptions of teachers regarding the integration of technology in STEM education. Investigating the motivations and challenges of STEM educators in underprivileged schools.
First is the peer pressure, to be followed by interest, abilities and family pressure. STEM strand can help students in their trainings for college through solving real world problems, critical thinking, and garner enough knowledge involving science and math that would be helpful in preparation in their chosen course.
55 Brilliant Research Topics For STEM Students Primarily, STEM is an acronym for Science, Technology, Engineering, and Mathematics. It's a study program that weaves all four disciplines for cross-disciplinary knowledge to solve scientific problems. STEM touches across a broad array of subjects as STEM students are required to gain mastery of four disciplines.
Use of a longitudinal study design allows for more precise analysis of the interplay between clonal hematopoiesis and atherosclerotic disease, finding that whereas clonal hematopoiesis confers an ...
Experimental design. We performed two observational studies (Fig. 1, Table 1) to study DCIS progression.In a cross-sectional study (Fig. 1A), we compared DCIS samples from patients with DCIS only (Pure DCIS, n = 58) versus DCIS samples from patients with synchronous invasive ductal carcinoma (Synchronous DCIS, n = 61).In a longitudinal case-control study (Fig. 1B), we collected samples from ...
For the research, Ruhl and her team studied 97 patients with sickle cell disease who underwent a low-intensity, or non-myeloablative, blood stem cell transplant between 2004-2019 at the NIH's ...